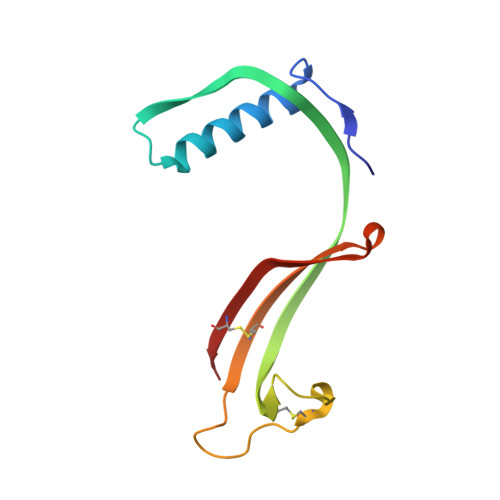
Domain swapping in N-truncated human cystatin C.
Janowski, R., Abrahamson, M., Grubb, A., Jaskolski, M.(2004) J Mol Biology 341: 151-160
- PubMed: 15312769 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.06.013
- Primary Citation Related Structures:
1R4C - PubMed Abstract:
Human cystatin C (HCC) inhibits papain-like cysteine proteases by a binding epitope composed of two beta-hairpin loops and the N-terminal segment. HCC is found in all body fluids and is present at a particularly high level in the cerebrospinal fluid. Oligomerization of HCC leads to amyloid deposits in brain arteries at advanced age but this pathological process is greatly accelerated with a naturally occurring Leu68Gln variant, resulting in fatal amyloidosis in early adult life. When proteins are extracted from human cystatin C amyloid deposits, an N-terminally truncated cystatin C (THCC) is found, lacking the first ten amino acid residues of the native sequence. It has been shown that the cerebrospinal fluid may cause this N-terminal truncation, possibly because of disintegration of the leucocytes normally present in this fluid, and the release of leucocyte proteolytic enzymes. HCC is the first disease-causing amyloidogenic protein for which oligomerization via 3D domain swapping has been observed. The aggregates arise in the crystallization buffer and have the form of 2-fold symmetric dimers in which a long alpha-helix of one molecule, flanked by two adjacent beta-strands, has replaced an identical domain of the other molecule, and vice versa. Consistent with a conformational change at one of the beta-hairpin loops of the binding epitope, the dimers (and also any other oligomers, including amyloid aggregates) are inactive as papain inhibitors. Here, we report the structure of N-truncated HCC, the dominant form of cystatin C in amyloid deposits. Although the protein crystallized under conditions that are drastically different from those for the full-length protein, the structure reveals dimerization by the same act of domain swapping. However, the new crystal structure is composed of four independent HCC dimers, none of which has the exact 2-fold symmetry of the full-length dimer. While the four dimers have the same overall topology, the exact relation between the individual domains shows a variability that reflects the flexibility at the dimer-specific open interface, which in the case of 3D domain-swapped HCC consists of beta-interactions between the open hinge loops and results in an unusually long intermolecular beta-sheet. The dimers are engaged in further quaternary interactions resulting in spherical, closed octameric assemblies that are identical to that present in the crystal of the full-length protein. The octamers interact via hydrophobic patches formed on the surface of the domain-swapped dimers as well as by extending the dimer beta-sheet through intermolecular contacts.
- Department of Crystallography, Faculty of Chemistry, A. Mickiewicz University, Grunwaldzka 6, 60-780 Poznan, Poland.
Organizational Affiliation: