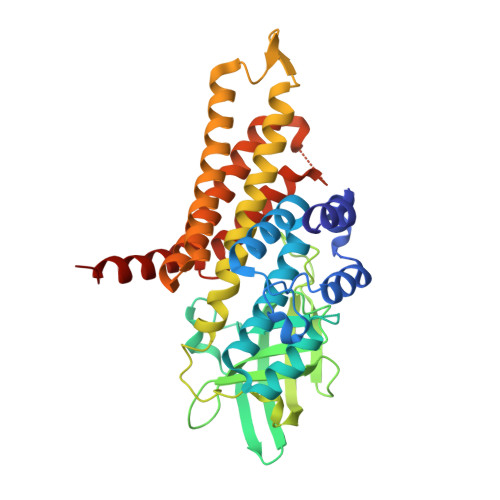
Crystal Structure of an Acyl-ACP Dehydrogenase from the FK520 Polyketide Biosynthetic Pathway: Insights into Extender Unit Biosynthesis
Watanabe, K., Khosla, C., Stroud, R.M., Tsai, S.-C.(2003) J Mol Biology 334: 435-444
- PubMed: 14623185 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.10.021
- Primary Citation Related Structures:
1R2J - PubMed Abstract:
Polyketide synthases (PKSs) synthesize the polyketide cores of pharmacologically important natural products such as the immunosuppressants FK520 and FK506. Understanding polyketide biosynthesis at atomic resolution could present new opportunities for chemo-enzymatic synthesis of complex molecules. The crystal structure of FkbI, an enzyme involved in the biosynthesis of the methoxymalonyl extender unit of FK520, was solved to 2.1A with an R(crys) of 24.4%. FkbI has a similar fold to acyl-CoA dehydrogenases. Notwithstanding this similarity, the surface and substrate-binding site of FkbI reveal key differences from other acyl-CoA dehydrogenases, suggesting that FkbI may recognize an acyl-ACP substrate rather than an acyl-CoA substrate. This structural observation coincided the genetic experiment done by Carroll et al. J. Am. Chem. Soc., 124 (2002) 4176. Although an in vitro assay for FkbI remains elusive, the structural basis for the substrate specificity of FkbI is analyzed by a combination of sequence comparison, docking simulations and structural analysis. A biochemical mechanism for the role of FkbI in the biosynthesis of methoxymalonyl-ACP is proposed.
- Department of Chemical Engineering, Stanford University, Stanford, CA 94305-5025, USA.
Organizational Affiliation: