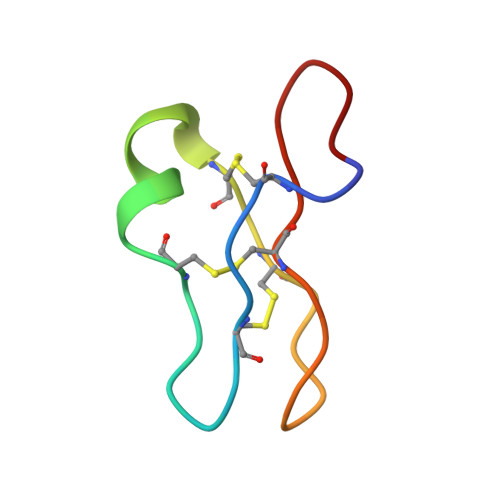
Solution structure of the cyclotide palicourein: implications for the development of a pharmaceutical framework.
Barry, D.G., Daly, N.L., Bokesch, H.R., Gustafson, K.R., Craik, D.J.(2004) Structure 12: 85-94
- PubMed: 14725768 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2003.11.019
- Primary Citation Related Structures:
1R1F - PubMed Abstract:
The cyclotides are a family of disulfide-rich proteins from plants. They have the characteristic structural features of a circular protein backbone and a knotted arrangement of disulfide bonds. Structural and biochemical studies of the cyclotides suggest that their unique physiological stability can be loaned to bioactive peptide fragments for pharmaceutical and agricultural development. In particular, the cyclotides incorporate a number of solvent-exposed loops that are potentially suitable for epitope grafting applications. Here, we determine the structure of the largest known cyclotide, palicourein, which has an atypical size and composition within one of the surface-exposed loops. The structural data show that an increase in size of a palicourein loop does not perturb the core fold, to which the thermodynamic and chemical stability has been attributed. The cyclotide core fold, thus, can in principle be used as a framework for the development of useful pharmaceutical and agricultural bioactivities.
- Institute for Molecular Bioscience, Queensland Bioscience Precinct, The University of Queensland Brisbane, Queensland 4072, Australia.
Organizational Affiliation: