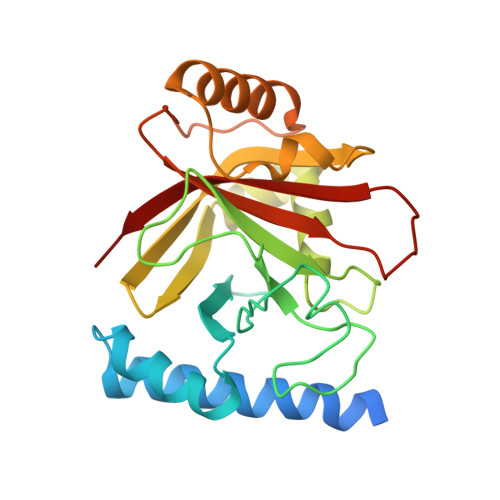
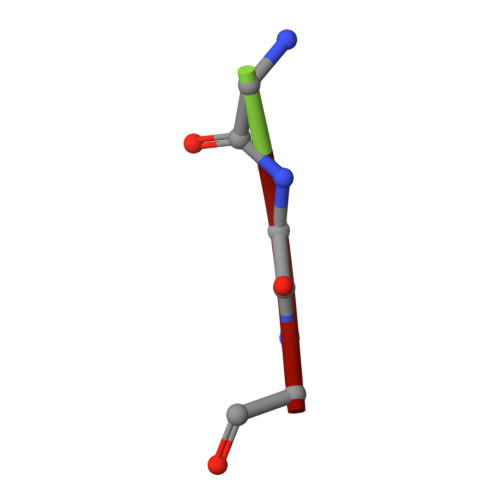
The structure of sortase B, a cysteine transpeptidase that tethers surface protein to the Staphylococcus aureus cell wall
Zong, Y., Mazmanian, S.K., Schneewind, O., Narayana, S.V.(2004) Structure 12: 105-112
- PubMed: 14725770 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2003.11.021
- Primary Citation Related Structures:
1QWZ, 1QX6, 1QXA - PubMed Abstract:
Many surface proteins of Gram-positive bacteria, which play important roles during the pathogenesis of human infections, are anchored to the cell wall envelope by a mechanism requiring sortases. Sortase B, a cysteine transpeptidase from Staphylococcus aureus, cleaves the C-terminal sorting signal of IsdC at the NPQTN motif and tethers the polypeptide to the pentaglycine cell wall cross-bridge. During catalysis, the active site cysteine of sortase and the cleaved substrate form an acyl intermediate, which is then resolved by the amino group of pentaglycine cross-bridges. We report here the crystal structures of SrtBDeltaN30 in complex with two active site inhibitors, MTSET and E64, and with the cell wall substrate analog tripleglycine. These structures reveal, for the first time, the active site disposition and the unique Cys-Arg catalytic machinery of the cysteine transpeptidase, and they also provide useful information for the future design of anti-infective agents against sortases.
- Center for Biophysical Sciences and Engineering, School of Optometry, University of Alabama at Birmingham, Birmingham, AL 35294, USA.
Organizational Affiliation: