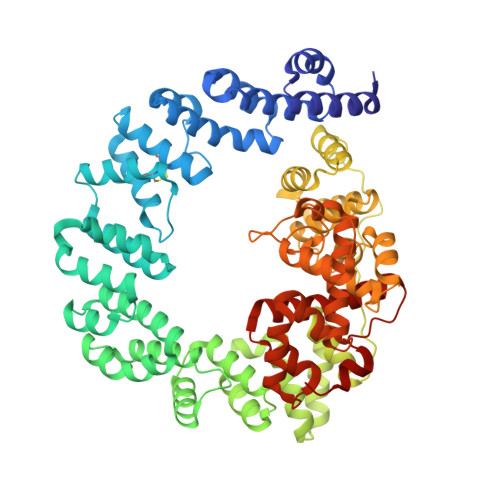
High resolution crystal structures of the Escherichia coli lytic transglycosylase Slt70 and its complex with a peptidoglycan fragment.
van Asselt, E.J., Thunnissen, A.M., Dijkstra, B.W.(1999) J Mol Biology 291: 877-898
- PubMed: 10452894 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.3013
- Primary Citation Related Structures:
1QSA, 1QTE - PubMed Abstract:
The 70 kDa soluble lytic transglycosylase (Slt70) from Escherichia coli is an exo-muramidase, that catalyses the cleavage of the glycosidic bonds between N -acetylmuramic acid and N -acetylglucosamine residues in peptidoglycan, the main structural component of the bacterial cell wall. This cleavage is accompanied by the formation of a 1,6-anhydro bond between the C1 and O6 atoms in the N -acetylmuramic acid residue (anhMurNAc). Crystallographic studies at medium resolution revealed that Slt70 is a multi-domain protein consisting of a large ring-shaped alpha-superhelix with on top a catalytic domain, which resembles the fold of goose-type lysozyme. Here we report the crystal structures of native Slt70 and of its complex with a 1,6-anhydromuropeptide solved at nominal resolutions of 1.65 A and 1.90 A, respectively. The high resolution native structure reveals the details on the hydrogen bonds, electrostatic and hydrophobic interactions that stabilise the catalytic domain and the alpha-superhelix. The building-block of the alpha-superhelix is an "up-down-up-down" four-alpha-helix bundle involving both parallel and antiparallel helix pairs. Stabilisation of the fold is provided through an extensive packing of apolar atoms, mostly from leucine and alanine residues. It lacks, however, an internal consensus sequence that characterises other super-secondary helical folds like the beta-helix in pectate lyase or the (beta-alpha)-helix in the ribonuclease inhibitor. The 1, 6-anhydromuropeptide product binds in a shallow groove adjacent to the peptidoglycan-binding groove of the catalytic domain. The groove is formed by conserved residues at the interface of the catalytic domain and the alpha-superhelix. The structure of the Slt70-1, 6-anhydromuropeptide complex confirms the presence of a specific binding-site for the peptide moieties of the peptidoglycan and it substantiates the notion that Slt70 starts the cleavage reaction at the anhMurNAc end of the peptidoglycan.
- University of Groningen, Nijenborgh 4, Groningen, 9747 AG, The Netherlands.
Organizational Affiliation: