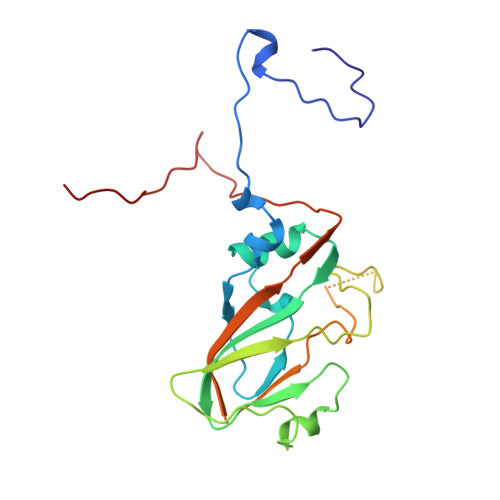
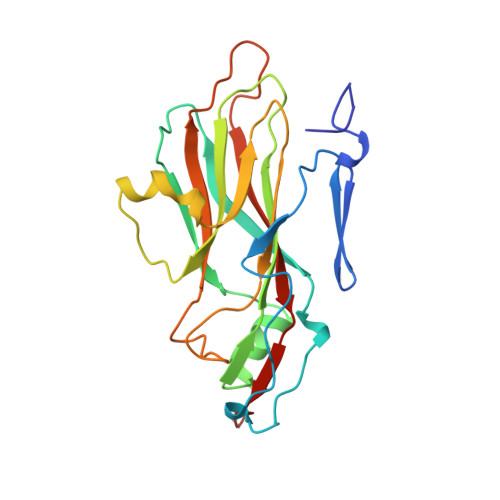
The structure and function of a foot-and-mouth disease virus-oligosaccharide receptor complex.
Fry, E.E., Lea, S.M., Jackson, T., Newman, J.W., Ellard, F.M., Blakemore, W.E., Abu-Ghazaleh, R., Samuel, A., King, A.M., Stuart, D.I.(1999) EMBO J 18: 543-554
- PubMed: 9927414 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/18.3.543
- Primary Citation Related Structures:
1QQP - PubMed Abstract:
Heparan sulfate has an important role in cell entry by foot-and-mouth disease virus (FMDV). We find that subtype O1 FMDV binds this glycosaminoglycan with a high affinity by immobilizing a specific highly abundant motif of sulfated sugars. The binding site is a shallow depression on the virion surface, located at the junction of the three major capsid proteins, VP1, VP2 and VP3. Two pre-formed sulfate-binding sites control receptor specificity. Residue 56 of VP3, an arginine in this virus, is critical to this recognition, forming a key component of both sites. This residue is a histidine in field isolates of the virus, switching to an arginine in adaptation to tissue culture, forming the high affinity heparan sulfate-binding site. We postulate that this site is a conserved feature of FMDVs, such that in the infected animal there is a biological advantage to low affinity, or more selective, interactions with glycosaminoglycan receptors.
- The Laboratory of Molecular Biophysics, Rex Richards Building, South Parks Road, Oxford OX1 3QU, UK.
Organizational Affiliation: