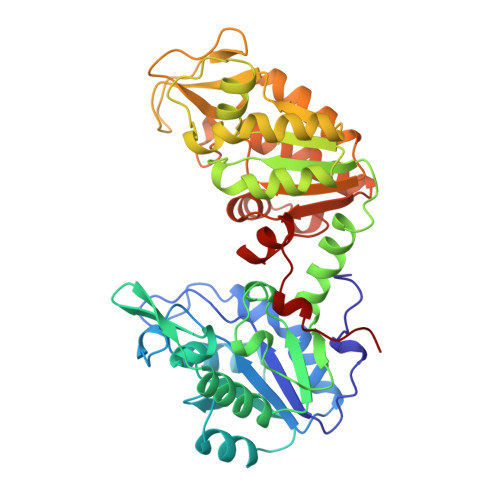
Structure of the R65Q mutant of yeast 3-phosphoglycerate kinase complexed with Mg-AMP-PNP and 3-phospho-D-glycerate.
McPhillips, T.M., Hsu, B.T., Sherman, M.A., Mas, M.T., Rees, D.C.(1996) Biochemistry 35: 4118-4127
- PubMed: 8672447 Search on PubMed
- DOI: https://doi.org/10.1021/bi952500o
- Primary Citation Related Structures:
1QPG - PubMed Abstract:
The structure of a ternary complex of the R65Q mutant of yeast 3-phosphoglycerate kinase (PGK) with magnesium 5'-adenylylimidodiphosphate (Mg-AMP-PNP) and 3-phospho-D-glycerate (3-PG) has been determined by X-ray crystallography to 2.4 angstrom resolution. The structure was solved by single isomorphous replacement, anamalous scattering, and solvent flattening and has been refined to an R-factor of 0.185, with rms deviations from ideal bond distance and angles of 0.009 angstrom and 1.78 degrees, respectively. PGK consists of two domains, with the 3-PG bound to a "basic patch" of residues from the N-terminal domain and the Mg-AMP-PNP interacting with residues from the C-terminal domain. The two ligands are separated by approximately 11 angstrom across the interdomain cleft. The model of the R65Q mutant of yeast PGK is very similar to the structures of PGK isolated from horse, pig, and Bacillus stearothermophilus (rms deviations between equivalent alpha-carbons in the individual domains < 1.0 angstrom) but exhibits substantial variations with a previously reported yeast structure (rms deviations between equivalent alpha-carbons in the individual domains of 2.9-3.2 angstrom). The most significant tertiary structural differences among the yeast R65Q, equine, porcine, and B. stearothermophilus PGK structures occur in the relative orientations of the two domains. However, the relationships between the observed conformations of PGK are inconsistent with a "hinge-bending" behavior that would close the interdomain cleft. It is proposed that the available structural and biochemical data on PGK may indicate that the basic patch primarily represents the site of anion activation and not the catalytically active binding site for 3-PG.
- Division of Chemistry, California Institute of Technology, Pasadena, 91125, USA.
Organizational Affiliation: