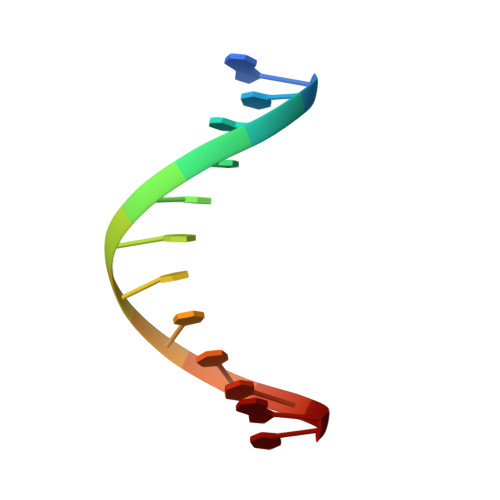
Solution Structure of the Complex between the Head-to-Tail Dimer of Calicheamicin Gamma-1-I Oligosaccharide and a DNA Duplex Containing D(ACCT) and D(TCCT) High-Affinity Binding Sites
Bifulco, G., Galeone, A., Nicolaou, K.C., Chazin, W.J., Gomez-Paloma, L.(1998) J Am Chem Soc 120: 7183-7191