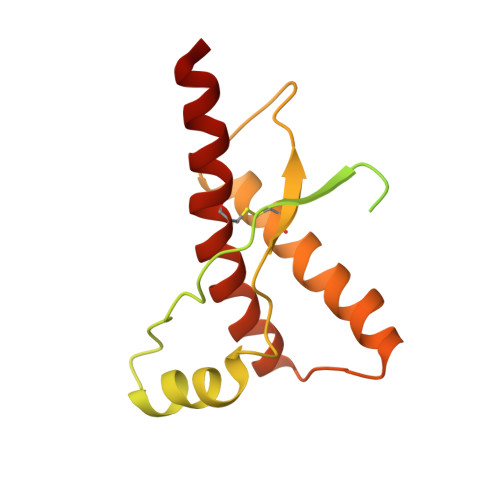
NMR Solution Structure of the Human Prion Protein
Zahn, R., Liu, A., Luhrs, T., Riek, R., Von Schroetter, C., Garcia, F.L., Billeter, M., Calzolai, L., Wider, G., Wuthrich, K.(2000) Proc Natl Acad Sci U S A 97: 145
- PubMed: 10618385 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.97.1.145
- Primary Citation Related Structures:
1QLX, 1QLZ, 1QM0, 1QM1, 1QM2, 1QM3 - PubMed Abstract:
The NMR structures of the recombinant human prion protein, hPrP(23-230), and two C-terminal fragments, hPrP(90-230) and hPrP(121-230), include a globular domain extending from residues 125-228, for which a detailed structure was obtained, and an N-terminal flexibly disordered "tail." The globular domain contains three alpha-helices comprising the residues 144-154, 173-194, and 200-228 and a short anti-parallel beta-sheet comprising the residues 128-131 and 161-164. Within the globular domain, three polypeptide segments show increased structural disorder: i.e., a loop of residues 167-171, the residues 187-194 at the end of helix 2, and the residues 219-228 in the C-terminal part of helix 3. The local conformational state of the polypeptide segments 187-193 in helix 2 and 219-226 in helix 3 is measurably influenced by the length of the N-terminal tail, with the helical states being most highly populated in hPrP(23-230). When compared with the previously reported structures of the murine and Syrian hamster prion proteins, the length of helix 3 coincides more closely with that in the Syrian hamster protein whereas the disordered loop 167-171 is shared with murine PrP. These species variations of local structure are in a surface area of the cellular form of PrP that has previously been implicated in intermolecular interactions related both to the species barrier for infectious transmission of prion disease and to immune reactions.
- Institut für Molekularbiologie und Biophysik, Eidgenössische Technische Hochschule Hönggerberg, CH-8093 Zürich, Switzerland.
Organizational Affiliation: