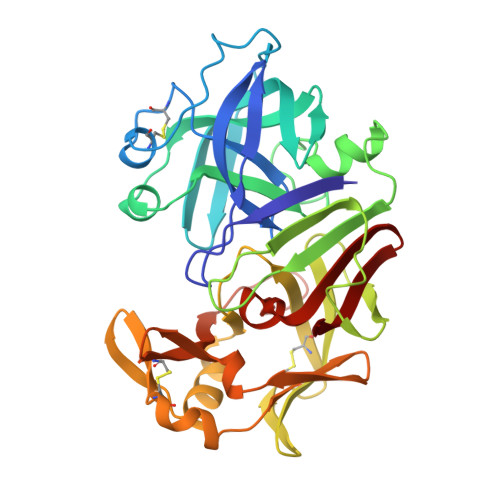
Structure of a pepsin/renin inhibitor complex reveals a novel crystal packing induced by minor chemical alterations in the inhibitor.
Chen, L., Erickson, J.W., Rydel, T.J., Park, C.H., Neidhart, D., Luly, J., Abad-Zapatero, C.(1992) Acta Crystallogr B 48: 476-488
- PubMed: 1418819 Search on PubMed
- DOI: https://doi.org/10.1107/s0108768192001939
- Primary Citation Related Structures:
1PSA - PubMed Abstract:
The structure determination by molecular replacement methods of a monoclinic pepsin/renin inhibitor complex crystal, with two molecules in the asymmetric unit, is presented. The atomic model, consisting of two liganded pepsin molecules and 110 water molecules, has been refined to a final crystallographic R value of 0.139 for data between 8 and 2.9 A resolution. The structure reveals a previously undescribed pepsin dimer formed predominantly by polar interactions. Inhibitor binding induces global structural changes in the native enzyme similar, but not identical, to the ones observed in other chemically similar pepsin/renin inhibitor complexes crystallized in an orthorhombic form. A region of the polypeptide chain (residues 292-297) which was not visible in the orthorhombic crystal is well ordered in the presently described structure; possibly induced by crystal contacts. The crystal packing of native pepsin is compared with the two different crystal forms of the inhibited enzyme.
- Laboratory of Protein Crystallography, Abbott Laboratories, Abbott Park, IL 60064.
Organizational Affiliation: