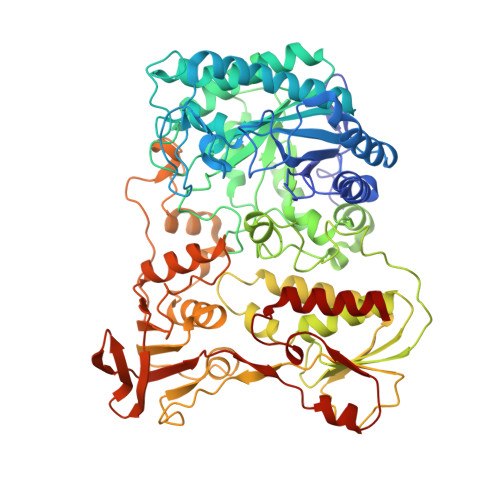
The crystal structure and reaction mechanism of Escherichia coli 2,4-dienoyl-CoA reductase
Hubbard, P.A., Liang, X., Schulz, H., Kim, J.J.(2003) J Biological Chem 278: 37553-37560
- PubMed: 12840019 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M304642200
- Primary Citation Related Structures:
1PS9 - PubMed Abstract:
Escherichia coli 2,4-dienoyl-CoA reductase is an iron-sulfur flavoenzyme required for the metabolism of unsaturated fatty acids with double bonds at even carbon positions. The enzyme contains FMN, FAD, and a 4Fe-4S cluster and exhibits sequence homology to another iron-sulfur flavoprotein, trimethylamine dehydrogenase. It also requires NADPH as an electron source, resulting in reduction of the C4-C5 double bond of the acyl chain of the CoA thioester substrate. The structure presented here of a ternary complex of E. coli 2,4-dienoyl-CoA reductase with NADP+ and a fatty acyl-CoA substrate reveals a possible mechanism for substrate reduction and provides details of a plausible electron transfer mechanism involving both flavins and the iron-sulfur cluster. The reaction is initiated by hydride transfer from NADPH to FAD, which in turn transfers electrons, one at a time, to FMN via the 4Fe-4S cluster. In the final stages of the reaction, the fully reduced FMN provides a hydride ion to the C5 atom of substrate, and Tyr-166 and His-252 are proposed to form a catalytic dyad that protonates the C4 atom of the substrate and complete the reaction. Inspection of the substrate binding pocket explains the relative promiscuity of the enzyme, catalyzing reduction of both 2-trans,4-cis- and 2-trans,4-trans-dienoyl-CoA thioesters.
- Department of Biochemistry, Medical College of Wisconsin, Milwaukee, Wisconsin 53226, USA.
Organizational Affiliation: