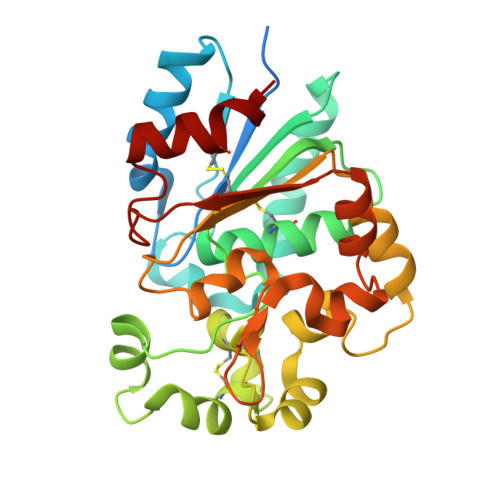
The crystal structure of palmitoyl protein thioesterase-2 (PPT2) reveals the basis for divergent substrate specificities of the two lysosomal thioesterases, PPT1 and PPT2.
Calero, G., Gupta, P., Nonato, M.C., Tandel, S., Biehl, E.R., Hofmann, S.L., Clardy, J.(2003) J Biological Chem 278: 37957-37964
- PubMed: 12855696 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M301225200
- Primary Citation Related Structures:
1PJA - PubMed Abstract:
Mutations in palmitoyl protein thioesterase-1 (PPT1) have been found to cause the infantile form of neuronal ceroid lipofuscinosis, which is a lysosomal storage disorder characterized by impaired degradation of fatty acid-modified proteins with accumulation of amorphous granular deposits in cortical neurons, leading to mental retardation and death. Palmitoyl protein thioesterase-2 (PPT2) is a second lysosomal hydrolase that shares a 26% identity with PPT1. A previous study had suggested that palmitoyl-CoA was the preferred substrate of PPT2. Furthermore, PPT2 did not hydrolyze palmitate from the several S-palmitoylated protein substrates. Interestingly, PPT2 deficiency in a recent transgenic mouse model is associated with a form of neuronal ceroid lipofuscinosis, suggesting that PPT1 and -2 perform non-redundant roles in lysosomal thioester catabolism. In the current paper, we present the crystal structure of PPT2 at a resolution of 2.7 A. Comparisons of the structures of PPT1 and -2 show very similar architectural features; however, conformational differences in helix alpha4 lead to a solvent-exposed lipid-binding groove in PPT1. The limited space between two parallel loops (beta3-alphaA and beta8-alphaF) located immediately above the lipid-binding groove in PPT2 restricts the binding of fatty acids with bulky head groups, and this binding groove is significantly larger in PPT1. This structural difference accounts for the ability of PPT2 to hydrolyze an unbranched structure such as palmitoyl-CoA but not palmitoylcysteine or palmitoylated proteins. Furthermore, differences in fatty acid chain length specificity of PPT1 and -2, also reported here, are explained by the structure and may provide a biochemical basis for their non-redundant roles.
- Department of Chemistry and Chemical Biology, Cornell University, Ithaca, New York 14853-1301, USA.
Organizational Affiliation: