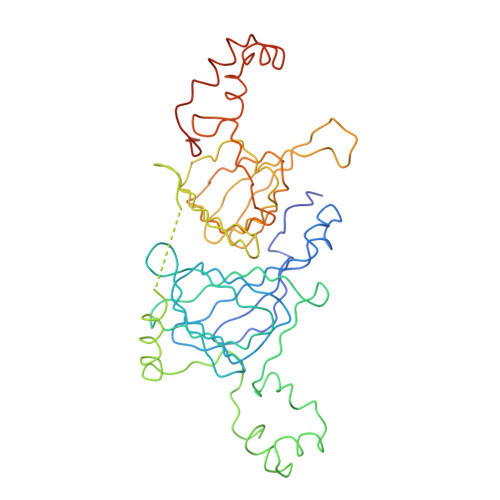
The three-dimensional structure of the seed storage protein phaseolin at 3 A resolution.
Lawrence, M.C., Suzuki, E., Varghese, J.N., Davis, P.C., Van Donkelaar, A., Tulloch, P.A., Colman, P.M.(1990) EMBO J 9: 9-15
- PubMed: 2295315 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/j.1460-2075.1990.tb08074.x
- Primary Citation Related Structures:
1PHS - PubMed Abstract:
The polypeptides of the trimeric seed storage protein phaseolin comprise two structurally similar units each made up of a beta-barrel and an alpha-helical domain. The beta-barrel has the 'jelly-roll' folding topology of the viral coat proteins and the alpha-helical domain shows structural similarity to the helix-turn-helix motif found in certain DNA-binding proteins.
- CSIRO Division of Biotechnology, Parkville, Victoria, Australia.
Organizational Affiliation: