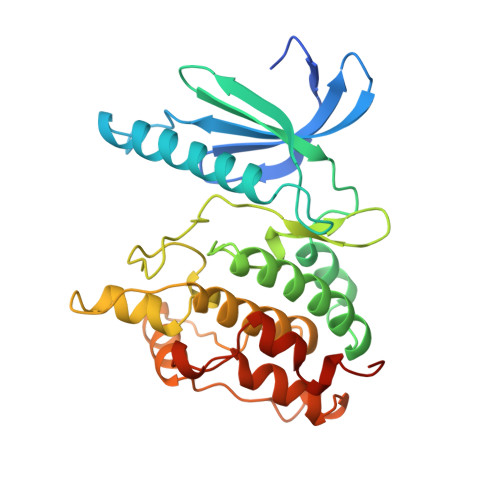
Two structures of the catalytic domain of phosphorylase kinase: an active protein kinase complexed with substrate analogue and product.
Owen, D.J., Noble, M.E., Garman, E.F., Papageorgiou, A.C., Johnson, L.N.(1995) Structure 3: 467-482
- PubMed: 7663944 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00180-0
- Primary Citation Related Structures:
1PHK - PubMed Abstract:
Control of intracellular events by protein phosphorylation is promoted by specific protein kinases. All the known protein kinase possess a common structure that defines a catalytically competent entity termed the 'kinase catalytic core'. Within this common structural framework each kinase displays its own unique substrate specificity, and a regulatory mechanism that may be modulated by association with other proteins. Structural studies of phosphorylase kinase (Phk), the major substrate of which is glycogen phosphorylase, may be expected to shed light on its regulation. We report two crystal structures of the catalytic core (residues 1-298; Phk gamma trnc) of the gamma-subunit of rabbit muscle phosphorylase kinase: the binary complex with Mn2+/beta-gamma-imidoadenosine 5'-triphosphate (AMPPNP) to a resolution of 2.6 A and the binary complex with Mg2+/ADP to a resolution of 3.0 A. The structures were solved by molecular replacement using the cAMP-dependent protein kinase (cAPK) as a model. The overall structure of Phk gamma trnc is similar to that of the catalytic core of other protein kinases. It consists of two domians joined on one edge by a 'hinge', with the catalytic site located in the cleft between the domains. Phk gamma trnc is constitutively active, and lacks the need for an activatory phosphorylation event that is essential for many kinases. The structure exhibits an essentially 'closed' conformation of the domains which is similar to that of cAPK complexed with substrates. The phosphorylated residue that is located at the domain interface in many protein kinases and that is believed to stabilize an active conformation is substituted by a glutamate in Phk gamma trnc. The glutamate, in a similar manner to the phosphorylated residue in other protein kinases, interacts with an arginine adjacent to the catalytic aspartate but does not participate in interdomain contacts. The interactions between the enzyme and the nucleotide product of its activity, Mg2+/ADP, explain the inhibitory properties of the nucleotides that are observed in kinetic studies.
- Laboratory of Molecular Biophysics, University of Oxford, UK.
Organizational Affiliation: