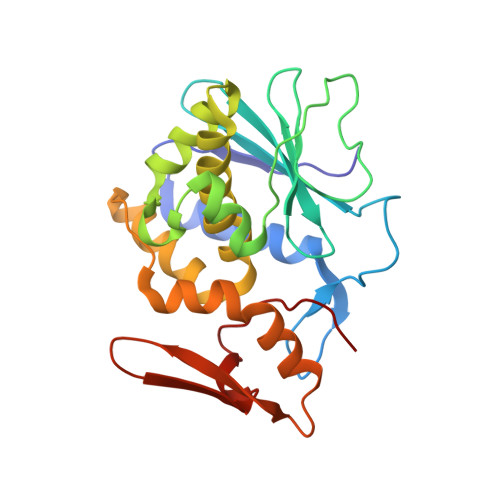
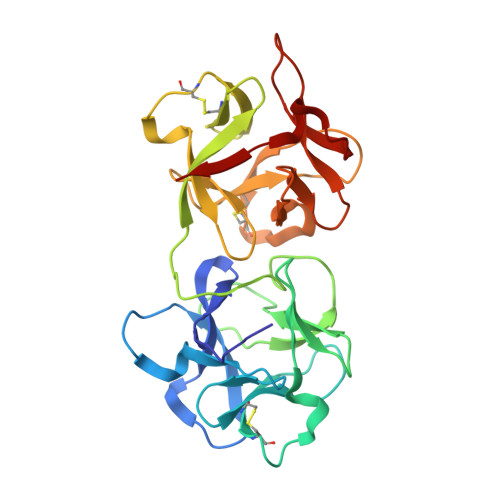
Structure of a novel ribosome-inactivating protein from a hemi-parasitic plant inhabiting the northwestern Himalayas.
Mishra, V., Ethayathulla, A.S., Sharma, R.S., Yadav, S., Krauspenhaar, R., Betzel, C., Babu, C.R., Singh, T.P.(2004) Acta Crystallogr D Biol Crystallogr 60: 2295-2304
- PubMed: 15583377 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904023534
- Primary Citation Related Structures:
1PC8 - PubMed Abstract:
This is the first report of the structural studies of a novel ribosome-inactivating protein (RIP) obtained from the Himalayan mistletoe (Viscum album) (HmRip). HmRip is a type II heterodimeric protein consisting of a toxic enzyme (A-chain) with an active site for ribosome inactivation and a lectin subunit (B-chain) with well defined sugar-binding sites. The crystal structure of HmRip has been determined at 3.8 A resolution and refined to a crystallographic R factor of 0.228 (R(free) = 0.271). A comparison of this structure with other type II RIPs reveals the presence of distinct structural features in the active site of the A-chain and in the 2gamma sugar-binding site of the B-chain. The conformation of the side chain of Tyr110, which is a conserved active-site residue in the A subunit, is strikingly different from those observed in other mistletoe RIPs, indicating its unique substrate-binding preference. The deletion of two important residues from the kink region after Ala231 in the 2gamma subdomain of the B-chain results in a significantly different conformation of the sugar-binding pocket. A ribosome-recognition site has also been identified in HmRip. The site is a shallow cavity, with the conserved residues Arg51, Asp70, Thr72 and Asn73 involved in the binding. The conformations of the antigenic epitopes of residues 1-20, 85-103 and 206-223 differ from those observed in other type II RIPs, resulting in the distinct antigenicity and pharmacological properties of HmRip.
- Department of Biophysics, All India Institute of Medical Sciences, New Delhi, India.
Organizational Affiliation: