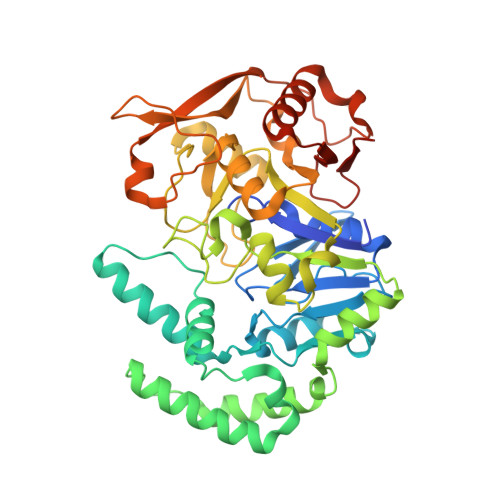
Crystal Structure of Fully Ligated Adenylosuccinate Synthetase from Plasmodium falciparum.
Eaazhisai, K., Jayalakshmi, R., Gayathri, P., Anand, R.P., Sumathy, K., Balaram, H., Murthy, M.R.(2004) J Mol Biology 335: 1251-1264
- PubMed: 14729341 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.11.036
- Primary Citation Related Structures:
1P9B - PubMed Abstract:
In the absence of the de novo purine nucleotide biosynthetic pathway in parasitic protozoa, purine salvage is of primary importance for parasite survival. Enzymes of the salvage pathway are, therefore, good targets for anti-parasitic drugs. Adenylosuccinate synthetase (AdSS), catalysing the first committed step in the synthesis of AMP from IMP, is a potential target for anti-protozoal chemotherapy. We report here the crystal structure of adenylosuccinate synthetase from the malaria parasite, Plasmodium falciparum, complexed to 6-phosphoryl IMP, GDP, Mg2+ and the aspartate analogue, hadacidin at 2 A resolution. The overall architecture of P. falciparum AdSS (PfAdSS) is similar to the known structures from Escherichia coli, mouse and plants. Differences in substrate interactions seen in this structure provide a plausible explanation for the kinetic differences between PfAdSS and the enzyme from other species. Additional hydrogen bonding interactions of the protein with GDP may account for the ordered binding of substrates to the enzyme. The dimer interface of PfAdSS is also different, with a pronounced excess of positively charged residues. Differences highlighted here provide a basis for the design of species-specific inhibitors of the enzyme.
- Molecular Biophysics Unit, UGC Centre of Advanced Study, Indian Institute of Science, Bangalore 560012, India.
Organizational Affiliation: