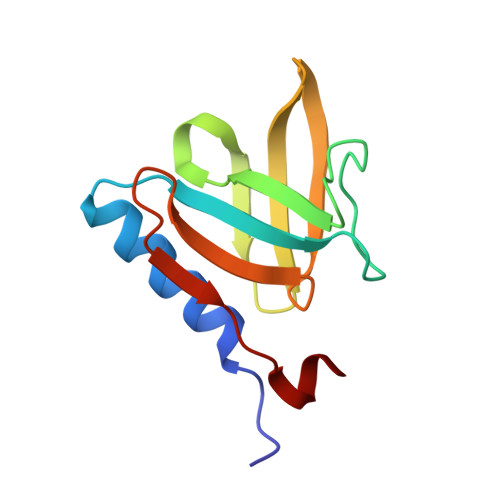
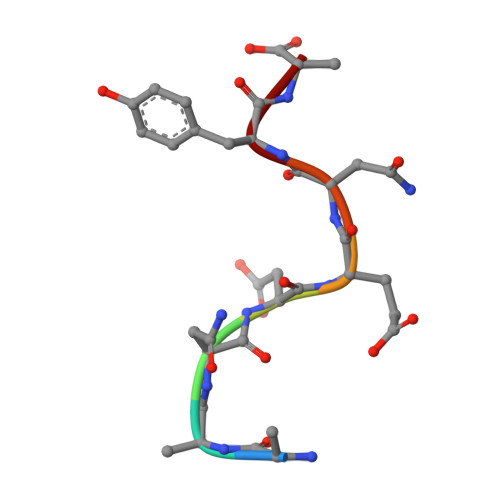
Structural basis of degradation signal recognition by SspB, a specificity-enhancing factor for the ClpXP proteolytic machine
Song, H.K., Eck, M.J.(2003) Mol Cell 12: 75-86
- PubMed: 12887894 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(03)00271-5
- Primary Citation Related Structures:
1OX8, 1OX9 - PubMed Abstract:
In prokaryotes, incomplete or misfolded polypeptides emanating from a stalled ribosome are marked for degradation by the addition of an 11 residue peptide (AANDENYALAA) to their C terminus. Substrates containing this conserved degradation signal, the SsrA tag, are targeted to specific proteases including ClpXP and ClpAP. SspB was originally characterized as a stringent starvation protein and has been found to bind specifically to SsrA-tagged proteins and to enhance recognition of these proteins by the ClpXP degradation machine. Here, we report the crystal structures of SspB alone and in complex with an SsrA peptide. Unexpectedly, SspB exhibits a fold found in Sm-family RNA binding proteins. The dimeric SspB structures explain the key determinants for recognition of the SsrA tag and define a hydrophobic channel that may bind unfolded substrates.
- Department of Cancer Biology, Dana-Farber Cancer Institute, 44 Binney Street, Boston, MA 02115, USA.
Organizational Affiliation: