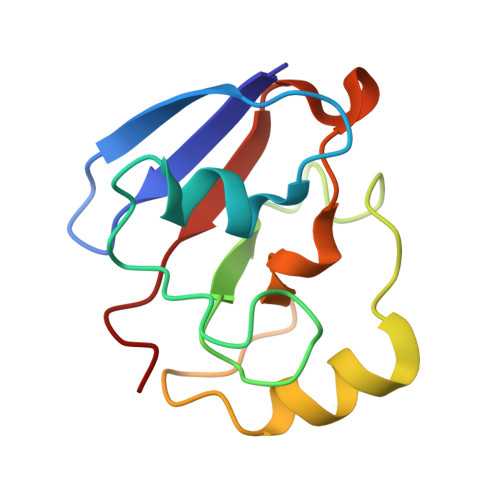
Crystal structure of putidaredoxin, the [2Fe-2S] component of the P450cam monooxygenase system from Pseudomonas putida
Sevrioukova, I.F., Garcia, C., Li, H., Bhaskar, B., Poulos, T.L.(2003) J Mol Biology 333: 377-392
- PubMed: 14529624 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.08.028
- Primary Citation Related Structures:
1OQQ, 1OQR - PubMed Abstract:
Stability of the [2Fe-2S]-containing putidaredoxin (Pdx), the electron donor to cytochrome P450cam in Pseudomonas putida, was improved by mutating non-ligating cysteine residues, Cys73 and Cys85, to serine singly and in combination. The increasing order of stability is Cys73Ser/Cys85Ser>Cys73Ser>Cys85Ser>WT Pdx. Crystal structures of Cys73Ser/Cys85Ser and Cys73Ser mutants of Pdx, solved by single-wavelength anomalous dispersion phasing using the [2Fe-2S] iron atoms to 1.47 A and 1.65 A resolution, respectively, are nearly identical and very similar to those of bovine adrenodoxin (Adx) and Escherichia coli ferredoxin. However, unlike the Adx structure, no motion between the core and interaction domains of Pdx is observed. This higher conformational stability of Pdx might be due to the presence of a more extensive hydrogen bonding network at the interface between the two structural domains around the conserved His49. In particular, formation of a hydrogen bond between the side-chain of Tyr51 and the carbonyl oxygen atom of Glu77 and the presence of two well-ordered water molecules linking the interaction domain and the C-terminal peptide to the core of the molecule are unique to Pdx. The folding topology of the NMR model is similar to that of the X-ray structure of Pdx. The overall rmsd of Calpha positions between the two models is 1.59 A. The largest positional differences are observed for residues 18-21 and 33-37 in the loop regions and the C terminus. The latter two peptides display conformational heterogeneity in the crystal structures. Owing to flexibility, the aromatic ring of the C-terminal Trp106 can closely approach the side-chains of Asp38 and Thr47 (3.2-3.9 A) or move away and leave the active site solvent-exposed. Therefore, Trp106, previously shown to be important in the Pdr-to-Pdx and Pdx-to-P450cam electron transfer reactions is in a position to regulate and/or mediate electron transfer to or from the [2Fe-2S] center of Pdx.
- Department of Molecular Biology and Biochemistry, University of California, Irvine, CA 92612-3900, USA. sevrioui@uci.edu
Organizational Affiliation: