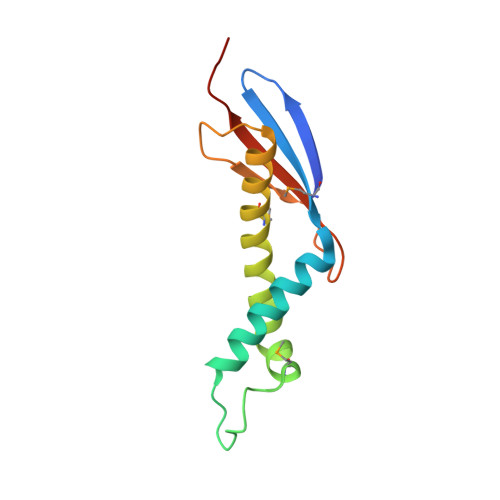
Chaperone binding at the ribosomal exit tunnel.
Kristensen, O., Gajhede, M.(2003) Structure 11: 1547-1556
- PubMed: 14656439 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2003.11.003
- Primary Citation Related Structures:
1OMS, 1P9Y - PubMed Abstract:
The exit tunnel region of the ribosome is well established as a focal point for interaction between the components that guide the fate of nascent polypeptides. One of these, the chaperone trigger factor (TF), associates with the 50S ribosomal subunit through its N-terminal domain. Targeting of TF to ribosomes is crucial to achieve its remarkable efficiency in protein folding. A similar tight coupling to translation is found in signal recognition particle (SRP)-dependent protein translocation. Here, we report crystal structures of the E. coli TF ribosome binding domain. TF is structurally related to the Hsp33 chaperone but has a prominent ribosome anchor located as a tip of the molecule. This tip includes the previously established unique TF signature motif. Comparison reveals that this feature is not found in SRP structures. We identify a conserved helical kink as a hallmark of the TF structure that is most likely critical to ensure ribosome association.
- Structural Biology, Department of Medicinal Chemistry, The Danish University of Pharmaceutical Sciences, Universitetsparken 2, DK-2100 Copenhagen, Denmark. ok@dfh.dk
Organizational Affiliation: