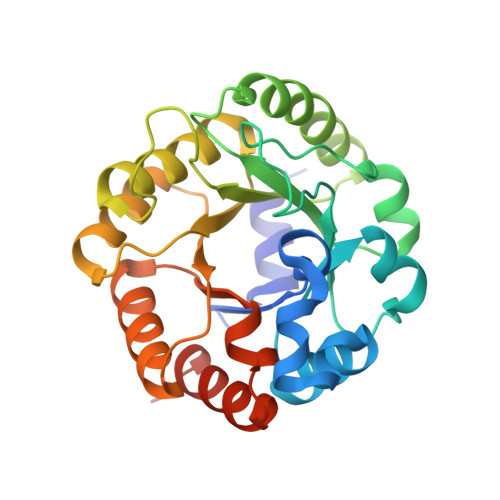
Crystal structure of an archaeal class I aldolase and the evolution of (betaalpha)8 barrel proteins.
Lorentzen, E., Pohl, E., Zwart, P., Stark, A., Russell, R.B., Knura, T., Hensel, R., Siebers, B.(2003) J Biological Chem 278: 47253-47260
- PubMed: 12941964 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M305922200
- Primary Citation Related Structures:
1OJX, 1OK4, 1OK6 - PubMed Abstract:
Fructose-1,6-bisphosphate aldolase (FBPA) catalyzes the reversible cleavage of fructose 1,6-bisphosphate to glyceraldehyde 3-phosphate and dihydroxyacetone phosphate in the glycolytic pathway. FBPAs from archaeal organisms have recently been identified and characterized as a divergent family of proteins. Here, we report the first crystal structure of an archaeal FBPA at 1.9-A resolution. The structure of this 280-kDa protein complex was determined using single wavelength anomalous dispersion followed by 10-fold non-crystallographic symmetry averaging and refined to an R-factor of 14.9% (Rfree 17.9%). The protein forms a dimer of pentamers, consisting of subunits adopting the ubiquitous (betaalpha)8 barrel fold. Additionally, a crystal structure of the archaeal FBPA covalently bound to dihydroxyacetone phosphate was solved at 2.1-A resolution. Comparison of the active site residues with those of classical FBPAs, which share no significant sequence identity but display the same overall fold, reveals a common ancestry between these two families of FBPAs. Structural comparisons, furthermore, establish an evolutionary link to the triosephosphate isomerases, a superfamily hitherto considered independent from the superfamily of aldolases.
- European Molecular Biology Laboratory, Hamburg Outstation, Notkestrasse 85, D-22603 Hamburg, Germany.
Organizational Affiliation: