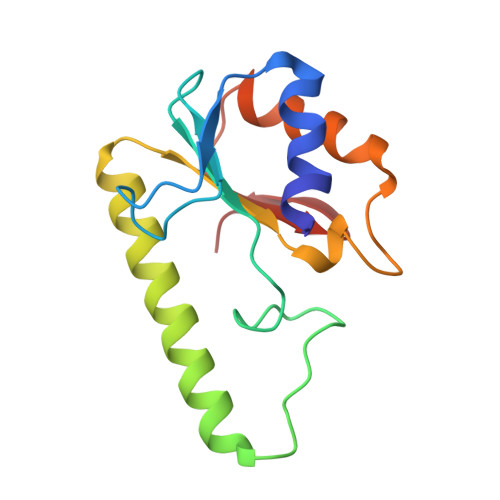
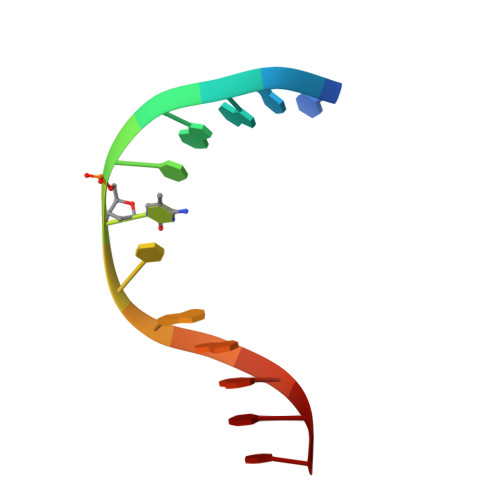
Crystal Structure of the Escherichia Coli Dcm Very-Short-Patch DNA Repair Endonuclease Bound to its Reaction Product-Site in a DNA Superhelix
Bunting, K.A., Roe, S.M., Headley, A., Brown, T., Savva, R., Pearl, L.H.(2003) Nucleic Acids Res 31: 1633
- PubMed: 12626704 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkg273
- Primary Citation Related Structures:
1ODG - PubMed Abstract:
Very-short-patch repair (Vsr) enzymes occur in a variety of bacteria, where they initiate nucleotide excision repair of G:T mismatches arising by deamination of 5-methyl-cytosines in specific regulatory sequences. We have now determined the structure of the archetypal dcm-Vsr endonuclease from Escherichia coli bound to the cleaved authentic hemi-deaminated/hemi-methylated dcm sequence 5'-C-OH-3' 5'-p-T-p-A-p-G-p-G-3'/3'-G-p-G-p-T-p(Me5)C-p-C formed by self-assembly of a 12mer oligonucleotide into a continuous nicked DNA superhelix. The structure reveals the presence of a Hoogsteen base pair within the deaminated recognition sequence and the substantial distortions of the DNA that accompany Vsr binding to product sites.
- Section of Structural Biology, Institute of Cancer Research, Chester Beatty Laboratories, 237 Fulham Road, London SW3 6JB, UK.
Organizational Affiliation: