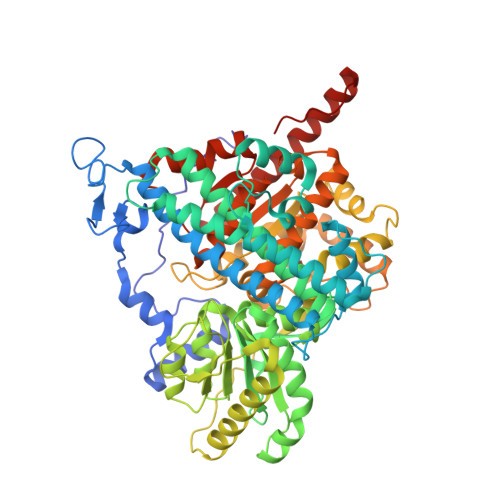
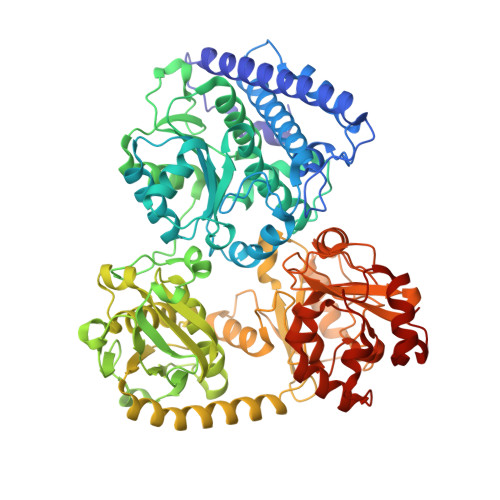
Ni-Zn-[Fe4-S4] and Ni-Ni-[Fe4-S4] Clusters in Closed and Open Alpha Subunits of Acetyl-Coa Synthase/Carbon Monoxide Dehydrogenase
Darnault, C., Volbeda, A., Kim, E.J., Legrand, P., Vernede, X., Lindahl, P.A., Fontecilla-Camps, J.C.(2003) Nat Struct Biol 10: 271
- PubMed: 12627225 Search on PubMed
- DOI: https://doi.org/10.1038/nsb912
- Primary Citation Related Structures:
1OAO - PubMed Abstract:
The crystal structure of the tetrameric alpha2beta2 acetyl-coenzyme A synthase/carbon monoxide dehydrogenase from Moorella thermoacetica has been solved at 1.9 A resolution. Surprisingly, the two alpha subunits display different (open and closed) conformations. Furthermore, X-ray data collected from crystals near the absorption edges of several metal ions indicate that the closed form contains one Zn and one Ni at its active site metal cluster (A-cluster) in the alpha subunit, whereas the open form has two Ni ions at the corresponding positions. Alternative metal contents at the active site have been observed in a recent structure of the same protein in which A-clusters contained one Cu and one Ni, and in reconstitution studies of a recombinant apo form of a related acetyl-CoA synthase. On the basis of our observations along with previously reported data, we postulate that only the A-clusters containing two Ni ions are catalytically active.
- Laboratoire de Cristallographie et Cristallogenèse des Protéines, Institut de Biologie Structurale 'Jean-Pierre Ebel', CEA, UJF, CNRS, 41, rue Jules Horowitz, 38027, Grenoble Cedex 1, France.
Organizational Affiliation: