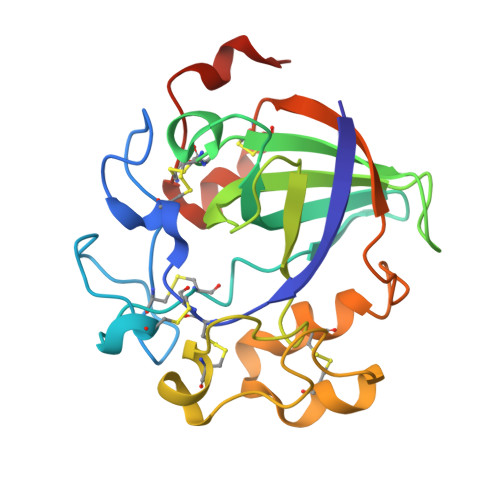
Crystal Structure of a Family 45 Endoglucanase from Melanocarpus Albomyces: Mechanistic Implications Based on the Free and Cellobiose-Bound Forms
Hirvonen, M., Papageorgiou, A.C.(2003) J Mol Biology 329: 403
- PubMed: 12767825 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00467-4
- Primary Citation Related Structures:
1OA7, 1OA9 - PubMed Abstract:
Cellulose, a polysaccharide of beta-1,4-linked D-glucosyl units, is the major component of plant cell walls and one of the most abundant biopolymers in nature. Cellulases (cellobiohydrolases and endoglucanases) are enzymes that catalyse the hydrolysis of cellulose to smaller oligosaccharides, a process of paramount importance in biotechnology. The thermophilic fungus Melanocarpus albomyces produces a 20 kDa endoglucanase known as 20K-cellulase that has been found particularly useful in the textile industry. The crystal structures of free 20K-cellulase and its complex with cellobiose have been determined at 2.0 A resolution. The enzyme, classified into the glycoside hydrolase family 45, exhibits the characteristic six-stranded beta-barrel found before in Humicola insolens endoglucanase V structure. However, the active site in the 20K-cellulase shows a closing of approximately 2.5-3.5A while a mobile loop identified previously in Humicola insolens endoglucanase V and implicated in the catalytic mechanism is well-defined in 20K-cellulase. In addition, the crystal structure of the cellobiose complex shows a shift in the cellobiose position at the substrate-binding cleft. It is therefore proposed that these alterations may reflect differences in the binding mechanism and catalytic action of the enzyme.
- Turku Centre for Biotechnology, University of Turku, P.O. Box 123, Turku 20521, Finland.
Organizational Affiliation: