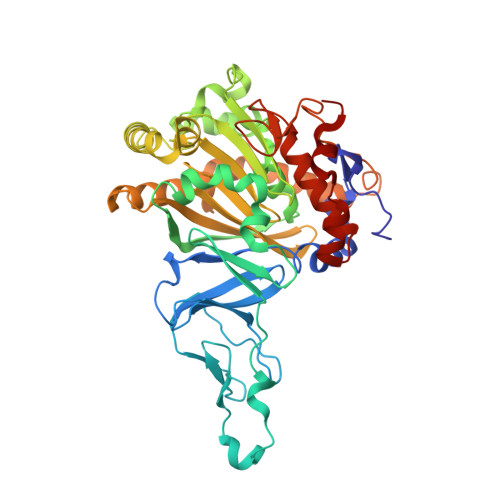
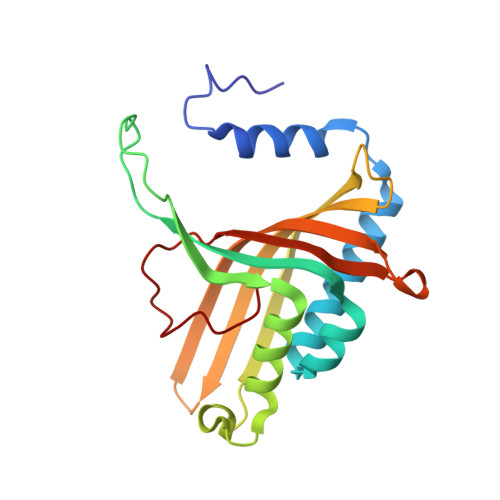
Crystal Structure of Naphthalene Dioxygenase: Side-on Binding of Dioxygen to Iron
Karlsson, A., Parales, J.V., Parales, R.E., Gibson, D.T., Eklund, H., Ramaswamy, S.(2003) Science 299: 1039
- PubMed: 12586937 Search on PubMed
- DOI: https://doi.org/10.1126/science.1078020
- Primary Citation Related Structures:
1O7G, 1O7H, 1O7M, 1O7N, 1O7P, 1O7W - PubMed Abstract:
Binding of oxygen to iron is exploited in several biological and chemical processes. Although computational and spectroscopic results have suggested side-on binding, only end-on binding of oxygen to iron has been observed in crystal structures. We have determined structures of naphthalene dioxygenase that show a molecular oxygen species bound to the mononuclear iron in a side-on fashion. In a complex with substrate and dioxygen, the dioxygen molecule is lined up for an attack on the double bond of the aromatic substrate. The structures reported here provide the basis for a reaction mechanism and for the high stereospecificity of the reaction catalyzed by naphthalene dioxygenase.
- Department of Molecular Biology, Swedish University of Agricultural Sciences, Box 590, Biomedical Center, 75124 Uppsala, Sweden.
Organizational Affiliation: