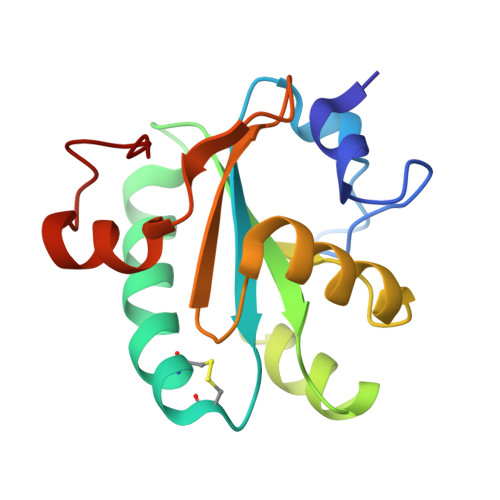
Tryparedoxins from Crithidia Fasciculata and Trypanosoma Brucei: Photoreduction of the Redox Disulfide Using Synchrotron Radiation and Evidence for a Conformational Switch Implicated in Function
Alphey, M.S., Gabrielsen, M., Micossi, E., Leonard, G.A., Mcsweeney, S.M., Ravelli, R.B.G., Tetaud, E., Fairlamb, A.H., Bond, C.S., Hunter, W.N.(2003) J Biological Chem 278: 25919
- PubMed: 12707277 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M301526200
- Primary Citation Related Structures:
1O73, 1O7U, 1O85, 1O8W, 1O8X, 1OC8, 1OC9 - PubMed Abstract:
Tryparedoxin (TryX) is a member of the thioredoxin (TrX) fold family involved in the regulation of oxidative stress in parasitic trypanosomatids. Like TrX, TryX carries a characteristic Trp-Cys-Xaa-Xaa-Cys motif, which positions a redox-active disulfide underneath a tryptophan lid. We report the structure of a Crithidia fasciculata tryparedoxin isoform (CfTryX2) in two crystal forms and compare them with structures determined previously. Efforts to chemically generate crystals of reduced TryX1 were unsuccessful, and we carried out a novel experiment to break the redox-active disulfide, formed between Cys-40 and Cys-43, utilizing the intense x-radiation from a third generation synchrotron undulator beamline. A time course study of the S-S bond cleavage is reported with the structure of a TryX1 C43A mutant as the control. When freed from the constraints of a disulfide link to Cys-43, Cys-40 pivots to become slightly more solvent-accessible. In addition, we have determined the structure of Trypanosoma brucei TryX, which, influenced by the molecular packing in the crystal lattice, displays a significantly different orientation of the active site tryptophan lid. This structural change may be of functional significance when TryX interacts with tryparedoxin peroxidase, the final protein in the trypanothione-dependent peroxidase pathway. Comparisons with chloroplast TrX and its substrate fructose 1,6-bisphosphate phosphatase suggest that this movement may represent a general feature of redox regulation in the trypanothione and thioredoxin peroxidase pathways.
- Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee, Dundee DD1 5EH, Scotland, United Kingdom.
Organizational Affiliation: