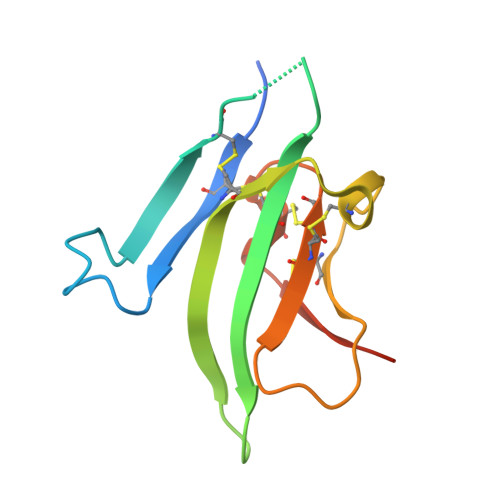
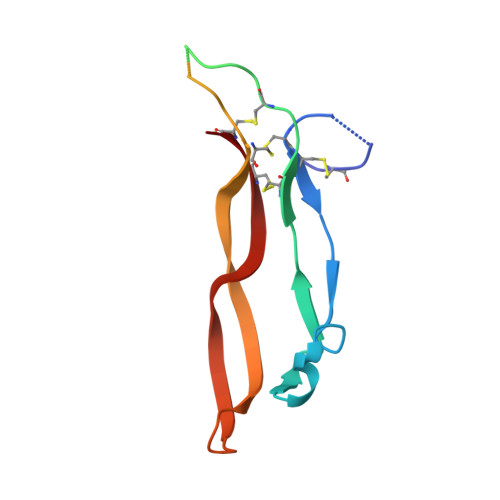
Structures of an ActRIIB:activin A complex reveal a novel binding mode for TGF-beta ligand:receptor interactions
Thompson, T.B., Woodruff, T.K., Jardetzky, T.S.(2003) EMBO J 22: 1555-1566
- PubMed: 12660162 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/cdg156
- Primary Citation Related Structures:
1NYS, 1NYU - PubMed Abstract:
The TGF-beta superfamily of ligands and receptors stimulate cellular events in diverse processes ranging from cell fate specification in development to immune suppression. Activins define a major subgroup of TGF-beta ligands that regulate cellular differentiation, proliferation, activation and apoptosis. Activins signal through complexes formed with type I and type II serine/threonine kinase receptors. We have solved the crystal structure of activin A bound to the extracellular domain of a type II receptor, ActRIIB, revealing the details of this interaction. ActRIIB binds to the outer edges of the activin finger regions, with the two receptors juxtaposed in close proximity, in a mode that differs from TGF-beta3 binding to type II receptors. The dimeric activin A structure differs from other known TGF-beta ligand structures, adopting a compact folded-back conformation. The crystal structure of the complex is consistent with recruitment of two type I receptors into a close packed arrangement at the cell surface and suggests that diversity in the conformational arrangements of TGF-beta ligand dimers could influence cellular signaling processes.
- Department of Biochemistry, Northwestern University, 2205 Tech Drive, Evanston, IL 60208, USA.
Organizational Affiliation: