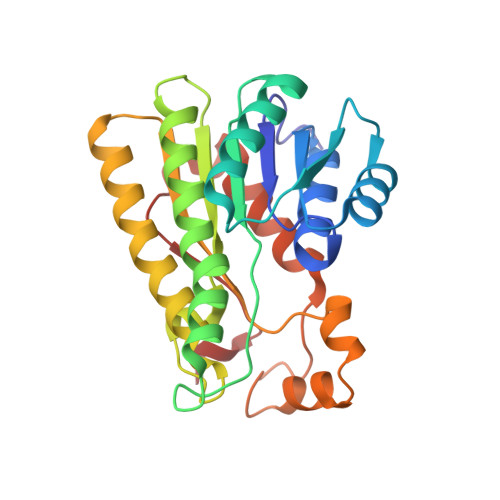
The crystal structure of R-specific alcohol dehydrogenase from Lactobacillus brevis suggests the structural basis of its metal dependency
Niefind, K., Muller, J., Riebel, B., Hummel, W., Schomburg, D.(2003) J Mol Biology 327: 317-328
- PubMed: 12628239 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00081-0
- Primary Citation Related Structures:
1NXQ - PubMed Abstract:
The crystal structure of the apo-form of an R-specific alcohol dehydrogenase from Lactobacillus brevis (LB-RADH) was solved and refined to 1.8A resolution. LB-RADH is a member of the short-chain dehydrogenase/reductase (SDR) enyzme superfamily. It is a homotetramer with 251 amino acid residues per subunit and uses NADP(H) as co-enzyme. NADPH and the substrate acetophenone were modelled into the active site. The enantiospecificity of the enzyme can be explained on the basis of the resulting hypothetical ternary complex. In contrast to most other SDR enzymes, the catalytic activity of LB-RADH depends strongly on the binding of Mg(2+). Mg(2+) removal by EDTA inactivates the enzyme completely. In the crystal structure, the Mg(2+)-binding site is well defined. The ion has a perfect octahedral coordination sphere and occupies a special position concerning crystallographic and molecular point symmetry, meaning that each RADH tetramer contains two magnesium ions. The magnesium ion is no direct catalytic cofactor. However, it is structurally coupled to the putative C-terminal hinge of the substrate-binding loop and, via an extended hydrogen bonding network, to some side-chains forming the substrate binding region. Therefore, the presented structure of apo-RADH provides plausible explanations for the metal dependence of the enzyme.
- Universität zu Köln, Institut für Biochemie, Zülpicher Strasse 47, Germany.
Organizational Affiliation: