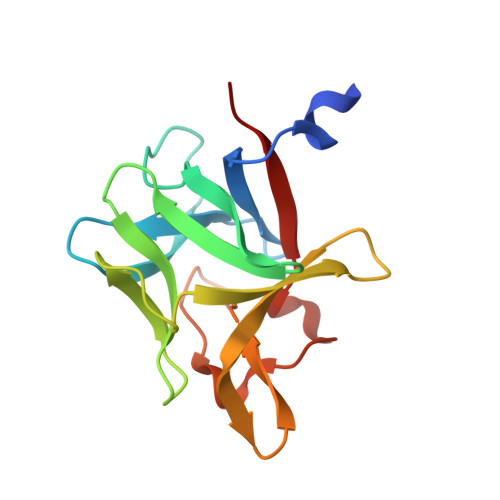
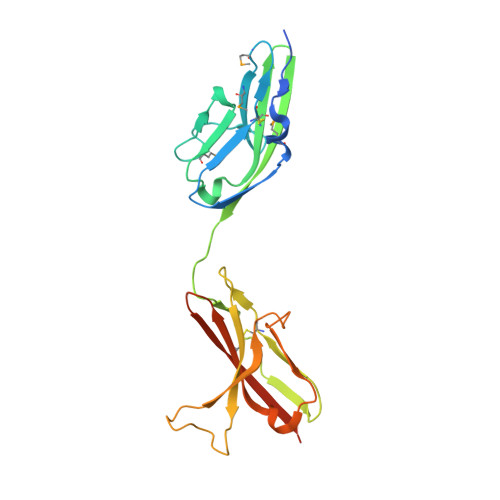
Structural basis by which alternative splicing confers specificity in fibroblast growth factor receptors.
Yeh, B.K., Igarashi, M., Eliseenkova, A.V., Plotnikov, A.N., Sher, I., Ron, D., Aaronson, S.A., Mohammadi, M.(2003) Proc Natl Acad Sci U S A 100: 2266-2271
- PubMed: 12591959 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0436500100
- Primary Citation Related Structures:
1NUN - PubMed Abstract:
Binding specificity between fibroblast growth factors (FGFs) and their receptors (FGFRs) is essential for mammalian development and is regulated primarily by two alternatively spliced exons, IIIb ("b") and IIIc ("c"), that encode the second half of Ig-like domain 3 (D3) of FGFRs. FGF7 and FGF10 activate only the b isoform of FGFR2 (FGFR2b). Here, we report the crystal structure of the ligand-binding portion of FGFR2b bound to FGF10. Unique contacts between divergent regions in FGF10 and two b-specific loops in D3 reveal the structural basis by which alternative splicing provides FGF10-FGFR2b specificity. Structure-based mutagenesis of FGF10 confirms the importance of the observed contacts for FGF10 biological activity. Interestingly, FGF10 binding induces a previously unobserved rotation of receptor Ig domain 2 (D2) to introduce specific contacts with FGF10. Hence, both D2 and D3 of FGFR2b contribute to the exceptional specificity between FGF10 and FGFR2b. We propose that ligand-induced conformational change in FGFRs may also play an important role in determining specificity for other FGF-FGFR complexes.
- Department of Pharmacology, New York University School of Medicine, New York, NY 10016, USA.
Organizational Affiliation: