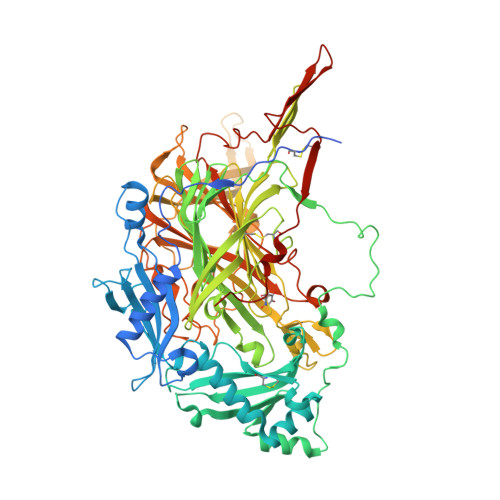
The Crystal Structure of Pichia pastoris Lysyl Oxidase
Duff, A.P., Cohen, A.E., Ellis, P.J., Kuchar, J.A., Langley, D.B., Shepard, E.M., Dooley, D.M., Freeman, H.C., Guss, J.M.(2003) Biochemistry 42: 15148-15157
- PubMed: 14690425 Search on PubMed
- DOI: https://doi.org/10.1021/bi035338v
- Primary Citation Related Structures:
1N9E - PubMed Abstract:
Pichia pastoris lysyl oxidase (PPLO) is unique among the structurally characterized copper amine oxidases in being able to oxidize the side chain of lysine residues in polypeptides. Remarkably, the yeast PPLO is nearly as effective in oxidizing a mammalian tropoelastin substrate as is a true mammalian lysyl oxidase isolated from bovine aorta. Thus, PPLO is functionally related to the copper-containing lysyl oxidases despite the lack of any significant sequence similarity with these enzymes. The structure of PPLO has been determined at 1.65 A resolution. PPLO is a homodimer in which each subunit contains a Type II copper atom and a topaquinone cofactor (TPQ) formed by the posttranslational modification of a tyrosine residue. While PPLO has tertiary and quaternary topologies similar to those found in other quinone-containing copper amine oxidases, its active site is substantially more exposed and accessible. The structural elements that are responsible for the accessibility of the active site are identified and discussed.
- School of Molecular and Microbial Biosciences, University of Sydney, Sydney, NSW, Australia.
Organizational Affiliation: