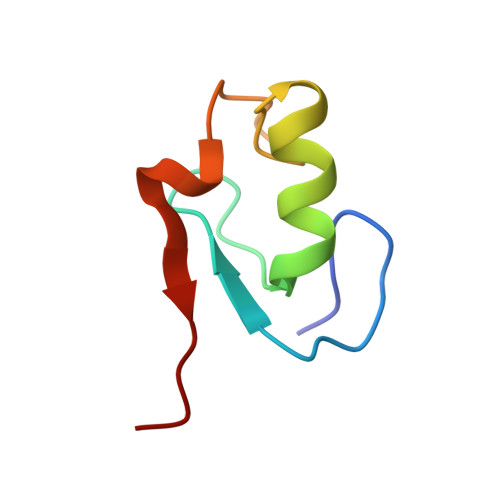
Structural insights into the U-box, a domain associated with multi-ubiquitination
Ohi, M.D., Vander Kooi, C.W., Rosenberg, J.A., Chazin, W.J., Gould, K.L.(2003) Nat Struct Biol 10: 250-255
- PubMed: 12627222 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsb906
- Primary Citation Related Structures:
1N87 - PubMed Abstract:
The structure of the U-box in the essential Saccharomyces cerevisiae pre-mRNA splicing factor Prp19p has been determined by NMR. The conserved zinc-binding sites supporting the cross-brace arrangement in RING-finger domains are replaced by hydrogen-bonding networks in the U-box. These hydrogen-bonding networks are necessary for the structural stabilization and activity of the U-box. A conservative Val-->Ile point mutation in the Prp19p U-box domain leads to pre-mRNA splicing defects in vivo. NMR analysis of this mutant shows that the substitution disrupts structural integrity of the U-box domain. Furthermore, comparison of the Prp19p U-box domain with known RING-E2 complex structures demonstrates that both U-box and RING-fingers contain a conserved interaction surface. Mutagenesis of residues at this interface, while not perturbing the structure of the U-box, abrogates Prp19p function in vivo. These comparative structural and functional analyses imply that the U-box and its associated ubiquitin ligase activity are critical for Prp19p function in vivo.
- Howard Hughes Medical Institute School of Medicine, Vanderbilt University, Nashville, Tennessee 37232, USA.
Organizational Affiliation: