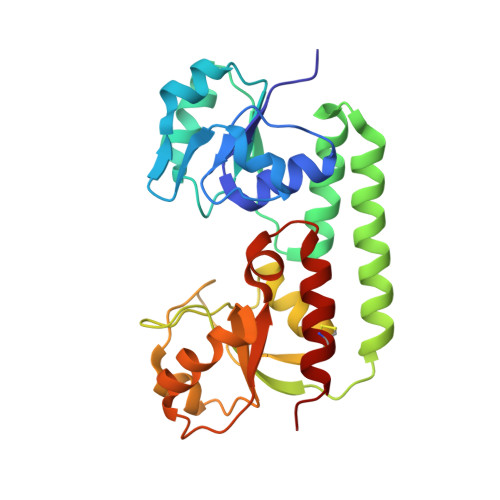
The structure of Escherichia coli BtuF and binding to its cognate ATP binding cassette transporter
Borths, E.L., Locher, K.P., Lee, A.T., Rees, D.C.(2002) Proc Natl Acad Sci U S A 99: 16642-16647
- PubMed: 12475936 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.262659699
- Primary Citation Related Structures:
1N2Z - PubMed Abstract:
Bacterial binding protein-dependent ATP binding cassette (ABC) transporters facilitate uptake of essential nutrients. The crystal structure of Escherichia coli BtuF, the protein that binds vitamin B12 and delivers it to the periplasmic surface of the ABC transporter BtuCD, reveals a bi-lobed fold resembling that of the ferrichrome binding protein FhuD. B12 is bound in the "base-on" conformation in a deep cleft formed at the interface between the two lobes of BtuF. A stable complex between BtuF and BtuCD (with the stoichiometry BtuC2D2F) is demonstrated to form in vitro and was modeled using the individual crystal structures. Two surface glutamates from BtuF may interact with arginine residues on the periplasmic surface of the BtuCD transporter. These glutamate and arginine residues are conserved among binding proteins and ABC transporters mediating iron and B12 uptake, suggesting that they may have a role in docking and the transmission of conformational changes.
- Howard Hughes Medical Institute and Division of Chemistry and Chemical Engineering, Mail Code 114-96, California Institute of Technology, Pasadena, CA 91125, USA.
Organizational Affiliation: