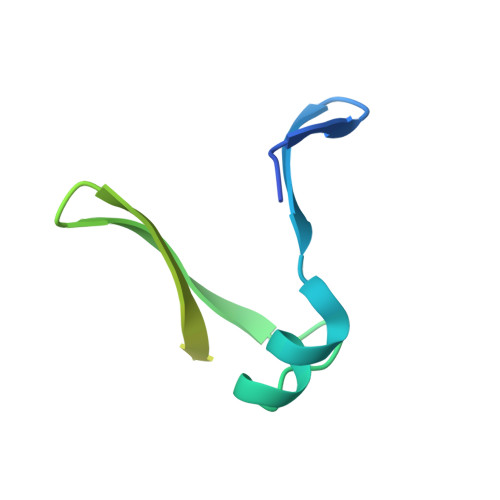
Crystal structure of the intrinsically flexible addiction antidote MazE.
Loris, R., Marianovsky, I., Lah, J., Laeremans, T., Engelberg-Kulka, H., Glaser, G., Muyldermans, S., Wyns, L.(2003) J Biological Chem 278: 28252-28257
- PubMed: 12743116 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M302336200
- Primary Citation Related Structures:
1MVF - PubMed Abstract:
A specific camel VHH (variable domain of dromedary heavy chain antibody) fragment was used to crystallize the intrinsically flexible addiction antidote MazE. Only 45% of the polypeptide chain is found ordered in the crystal. The MazE monomer consisting of two beta-hairpins connected by a short alpha-helix has no hydrophobic core on its own and represents only one half of a typical protein domain. A complete domain structure is formed by the association of two chains, creating a hydrophobic core between two four-stranded beta-sheets. This hydrophobic core consists exclusively of short aliphatic residues. The folded part of MazE contains a novel DNA binding motif. A model for DNA binding that is consistent with the available biochemical data is presented.
- Laboratorium voor Ultrastructuur en Vlaams instituut voor Biotechnologie, Vrije Universiteit Brussel, Gebouw E, Pleinlaan 2, 1050 Brussel, Belgium. reloris@vub.ac.be
Organizational Affiliation: