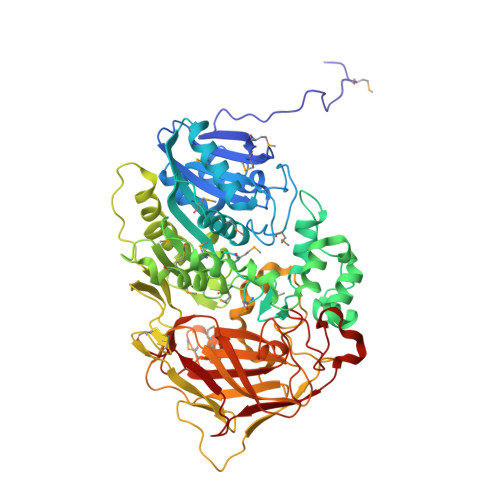
The sequence and crystal structure of the alpha-amino acid ester hydrolase from Xanthomonas citri define a new family of beta-lactam antibiotic acylases.
Barends, T.R., Polderman-Tijmes, J.J., Jekel, P.A., Hensgens, C.M., de Vries, E.J., Janssen, D.B., Dijkstra, B.W.(2003) J Biological Chem 278: 23076-23084
- PubMed: 12684501 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M302246200
- Primary Citation Related Structures:
1MPX - PubMed Abstract:
alpha-Amino acid ester hydrolases (AEHs) catalyze the hydrolysis and synthesis of esters and amides with an alpha-amino group. As such, they can synthesize beta-lactam antibiotics from acyl compounds and beta-lactam nuclei obtained from the hydrolysis of natural antibiotics. This article describes the gene sequence and the 1.9-A resolution crystal structure of the AEH from Xanthomonas citri. The enzyme consists of an alpha/beta-hydrolase fold domain, a helical cap domain, and a jellyroll beta-domain. Structural homology was observed to the Rhodococcus cocaine esterase, indicating that both enzymes belong to the same class of bacterial hydrolases. Docking of a beta-lactam antibiotic in the active site explains the substrate specificity, specifically the necessity of an alpha-amino group on the substrate, and explains the low specificity toward the beta-lactam nucleus.
- Department of Biophysical Chemistry, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, Nijenborgh 4, NL-9747 AG Groningen, The Netherlands.
Organizational Affiliation: