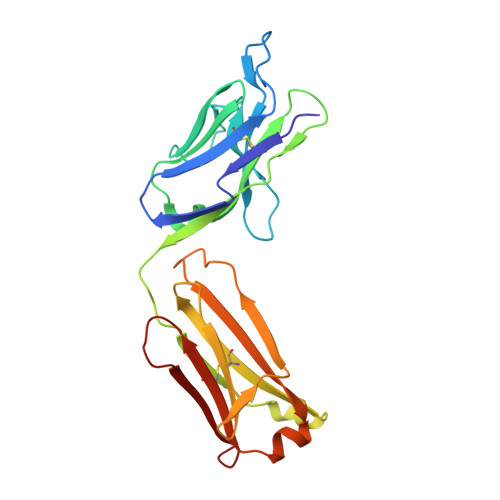
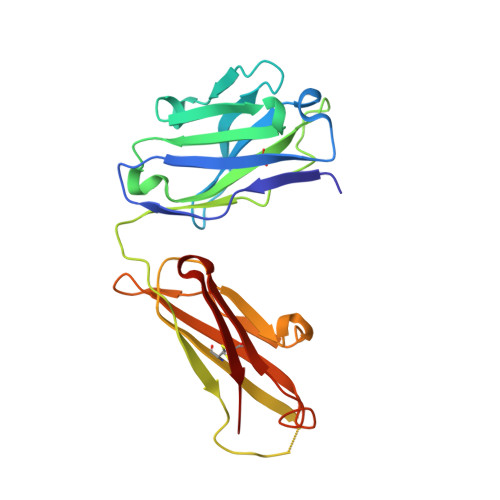
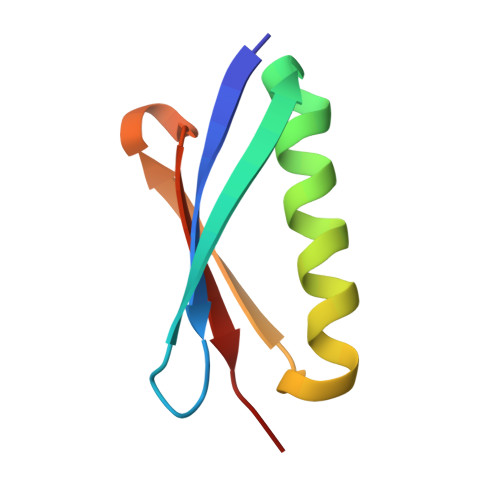
Evidence for plasticity and structural mimicry at the immunoglobulin light chain-protein L interface
Graille, M., Harrison, S., Crump, M.P., Findlow, S.C., Housden, N.G., Muller, B.H., Battail-Poirot, N., Sibai, G., Sutton, B.J., Taussig, M.J., Jolivet-Reynaud, C., Gore, M.G., Stura, E.A.(2002) J Biological Chem 277: 47500-47506
- PubMed: 12221088 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M206105200
- Primary Citation Related Structures:
1MHH - PubMed Abstract:
The multidomain bacterial surface protein L (PpL) is a virulence factor expressed by only 10% of Peptostreptococcus magnus strains, and its expression is correlated with bacterial vaginosis. The molecular basis for its ability to recognize 60% of mammalian immunoglobulin light chain variable regions (V(L)) has been described recently by x-ray crystallography, which suggested the presence of two V(L) binding sites on each protein L domain (Graille, M., Stura, E. A., Housden, N. G., Beckingham, J. A., Bottomley, S. P., Beale, D., Taussig, M. J., Sutton, B. J., Gore, M. G., and Charbonnier, J. (2001) Structure 9, 679-687). Here, we report the crystal structure at 2.1 A resolution of a protein L mutant complexed to an Fab' fragment with only 50% of the V(L) residues interacting with PpL site 1 conserved. Comparison of the site 1 interface from both structures shows how protein L is able to accommodate these sequence differences and therefore bind to a large repertoire of Ig. The x-ray structure and NMR results confirm the existence of two V(L) binding sites on a single protein L domain. These sites exhibit a remarkable structural mimicry of growth factors binding to their receptors. This could explain the protein L superantigenic activity on human B lymphocytes.
- Laboratoire de Structure des Protéines, Département d'Ingénierie et d'Etudes des Protéines (DIEP), Commissariat à l'Energie Atomique, Centre d'Etudes de Saclay, Gif-sur-Yvette, France.
Organizational Affiliation: