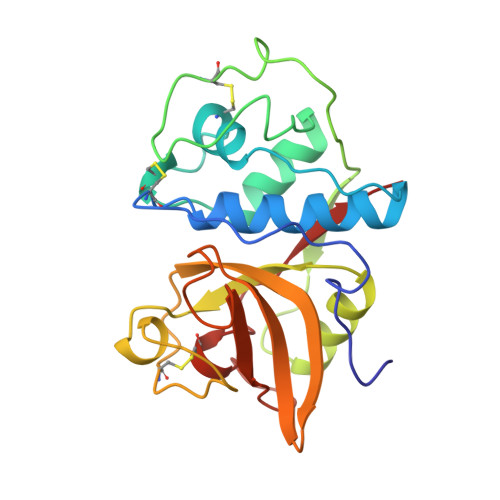
Crystal structure of a caricain D158E mutant in complex with E-64.
Katerelos, N.A., Taylor, M.A., Scott, M., Goodenough, P.W., Pickersgill, R.W.(1996) FEBS Lett 392: 35-39
- PubMed: 8769310 Search on PubMed
- DOI: https://doi.org/10.1016/0014-5793(96)00697-7
- Primary Citation Related Structures:
1MEG - PubMed Abstract:
The structure of the D158E mutant of caricain (previously known as papaya protease omega) in complex with E-64 has been determined at 2.0 A resolution (overall R factor 19.3%). The structure reveals that the substituted glutamate makes the same pattern of hydrogen bonds as the aspartate in native caricain. This was not anticipated since in the native structure there is insufficient room to accommodate the glutamate side chain. The glutamate is accommodated in the mutant by a local expansion of the structure demonstrating that small structural changes are responsible for the change in activity.
- Institute of Food Research, Reading Laboratory, UK.
Organizational Affiliation: