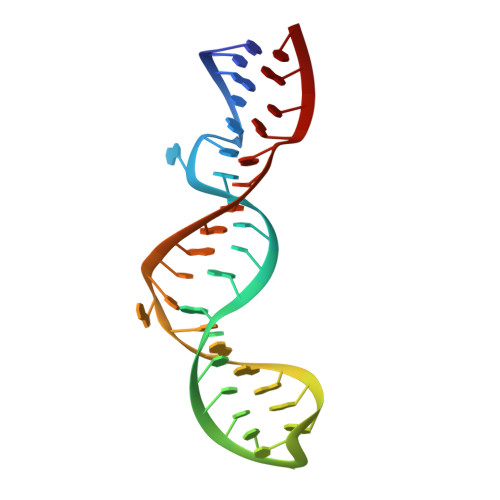
Structure and stability of wild-type and mutant RNA internal loops from the SL-1 domain of the HIV-1 packaging signal
Greatorex, J., Gallego, J., Varani, G., Lever, A.(2002) J Mol Biology 322: 543-557
- PubMed: 12225748 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)00776-3
- Primary Citation Related Structures:
1M5L - PubMed Abstract:
The packaging signal (Psi) of the human immunodeficiency virus type 1 (HIV-1) enables encapsidation of the full-length genomic RNA against a background of a vast excess of cellular mRNAs. The core HIV-1 Psi is approximately 109 nucleotides and contains sequences critical for viral genomic dimerisation and splicing, in addition to the packaging signal. It consists of a series of stem-loops (termed SL-1 to SL-4), which can be arranged in a cloverleaf secondary structure. Using a combination of NMR spectroscopy, UV melting experiments, molecular modeling and phylogenetic analyses, we have explored the structure of two conserved internal loops proximal to the palindromic sequence of SL-1. Internal loop A, composed of six purines, forms a flexible structure that is strikingly similar to the Rev responsive element motif when bound to Rev protein. This result suggests that it may function as a protein-binding site. The absolutely conserved four-purine internal loop B is instead conformationally and thermodynamically unstable, and exhibits multiple conformations in solution. By introducing a double AGG to GGA mutation within this loop, its conformation is stabilised to form a new intra-molecular G:A:G base-triplet. The structure of the GGA mutant explains the relative instability of the wild-type loop. In a manner analogous to SL-3, we propose that conformational flexibility at this site may facilitate melting of the structure during Gag protein capture or genomic RNA dimerisation.
- Department of Medicine, University of Cambridge, Level 5, Addenbrookes Hospital, Hills Road, CB2 2QQ, Cambridge, UK.
Organizational Affiliation: