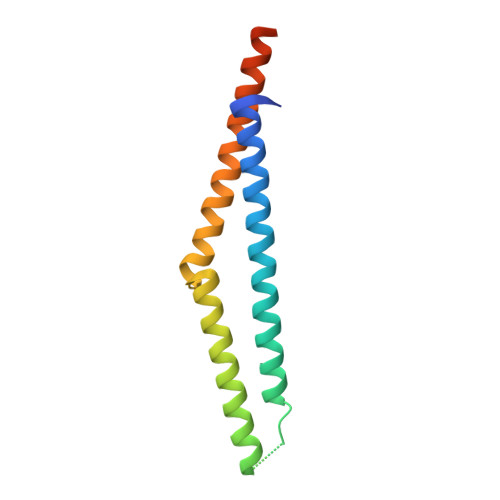
The coiled coil region (amino acids 129-250) of the tumor suppressor protein adenomatous polyposis coli (APC). Its structure and its interaction with chromosome maintenance region 1 (Crm-1).
Tickenbrock, L., Cramer, J., Vetter, I.R., Muller, O.(2002) J Biological Chem 277: 32332-32338
- PubMed: 12070164 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M203990200
- Primary Citation Related Structures:
1M5I - PubMed Abstract:
The APC (adenomatous polyposis coli) tumor suppressor protein has many different intracellular functions including a nuclear export activity. Only little is known about the molecular architecture of the 2843-amino acid APC protein. Guided by secondary structure predictions we identified a fragment close to the N-terminal end, termed APC-(129-250), as a soluble and protease-resistant domain. We solved the crystal structure of APC-(129-250), which is monomeric and consists of three alpha-helices forming two separate antiparallel coiled coils. APC-(129-250) includes the nuclear export signal NES-(165-174) at the C-terminal end of the first helix. Surprisingly, the conserved hydrophobic amino acids of NES-(165-174) are buried in one of the coiled coils and are thus not accessible for interaction with other proteins. We demonstrate the direct interaction of APC-(129-250) with the nuclear export factor chromosome maintenance region 1 (Crm-1). This interaction is enhanced by the small GTPase Ran in its activated GTP-bound form and also by a double mutation in APC-(129-250), which deletes two amino acids forming two of the major interhelical interactions within the coiled coil. These observations hint to a regulatory mechanism of the APC nuclear export activity by NES masking.
- Max-Planck-Institut für molekulare Physiologie, Abteilung Strukturelle Biologie, Otto-Hahn-Strasse 11, D-44227 Dortmund, Germany.
Organizational Affiliation: