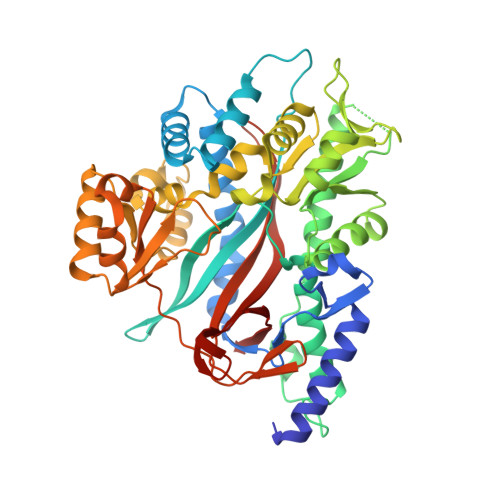
Large Conformational Changes in the Catalytic Cycle of Glutathione Synthase
Gogos, A., Shapiro, L.(2002) Structure 10: 1669-1676
- PubMed: 12467574 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00906-1
- Primary Citation Related Structures:
1M0T, 1M0W - PubMed Abstract:
Glutathione synthase catalyzes the final ATP-dependent step in glutathione biosynthesis, the formation of glutathione from gamma-glutamylcysteine and glycine. We have determined structures of yeast glutathione synthase in two forms: unbound (2.3 A resolution) and bound to its substrate gamma-glutamylcysteine, the ATP analog AMP-PNP, and two magnesium ions (1.8 A resolution). These structures reveal that upon substrate binding, large domain motions convert the enzyme from an open unliganded form to a closed conformation in which protein domains completely surround the substrate in the active site.
- Department of Biochemistry and Molecular Biophysics, Columbia University College of Physicians and Surgeons, 630 West 168th Street, New York, NY 10032, USA.
Organizational Affiliation: