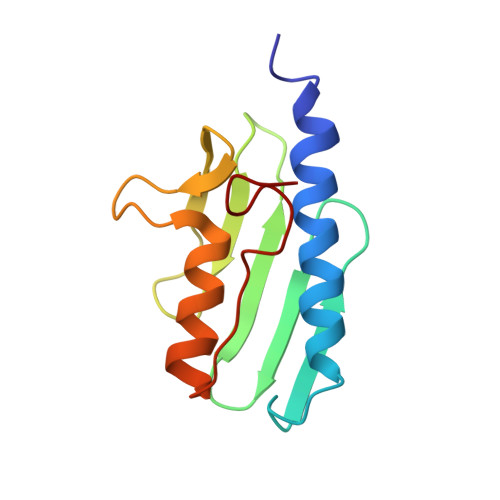
Towards a structural understanding of Friedreich's ataxia: the solution structure of frataxin
Musco, G., Stier, G., Kolmerer, B., Adinolfi, S., Martin, S., Frenkiel, T., Gibson, T., Pastore, A.(2000) Structure 8: 695-707
- PubMed: 10903947 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(00)00158-1
- Primary Citation Related Structures:
1LY7 - PubMed Abstract:
Lesions in the gene for frataxin, a nuclear-encoded mitochondrial protein, cause the recessively inherited condition Friedreich's ataxia. It is thought that the condition arises from disregulation of mitochondrial iron homeostasis, with concomitant oxidative damage leading to neuronal death. Very little is, as yet, known about the biochemical function of frataxin. Here, we show that the mature form of recombinant frataxin behaves in solution as a monodisperse species that is composed of a 15-residue-long unstructured N terminus and an evolutionarily conserved C-terminal region that is able to fold independently. The structure of the C-terminal domain consists of a stable seven-stranded antiparallel beta sheet packing against a pair of parallel helices. The structure is compact with neither grooves nor cavities, features that are typical of iron-binding modules. Exposed evolutionarily conserved residues cover a broad area and all cluster on the beta-sheet face of the structure, suggesting that this is a functionally important surface. The effect of two clinically occurring mutations on the fold was checked experimentally. When the mature protein was titrated with iron, no tendency to iron-binding or to aggregation was observed. Knowledge of the frataxin structure provides important guidelines as to the nature of the frataxin binding partner. The absence of all the features expected for an iron-binding activity, the large conserved area on its surface and lack of evidence for iron-binding activity strongly support an indirect involvement of frataxin in iron metabolism. The effects of point mutations associated with Friedreich's ataxia can be rationalised by knowledge of the structure and suggest possible models for the occurrence of the disease in compound heterozygous patients.
- NIMR, Mill Hill, NW7 1AA, UK.
Organizational Affiliation: