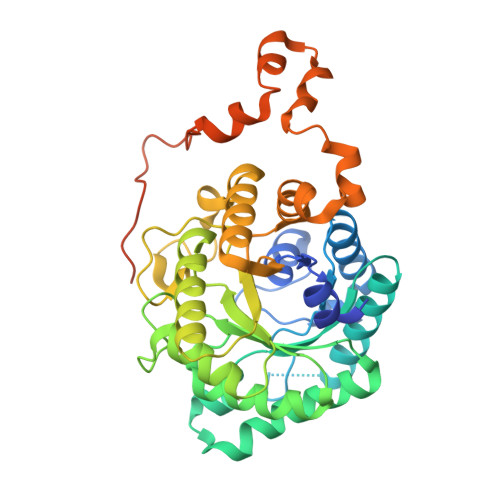
Betaine-homocysteine methyltransferase: zinc in a distorted barrel.
Evans, J.C., Huddler, D.P., Jiracek, J., Castro, C., Millian, N.S., Garrow, T.A., Ludwig, M.L.(2002) Structure 10: 1159-1171
- PubMed: 12220488 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00796-7
- Primary Citation Related Structures:
1LT7, 1LT8 - PubMed Abstract:
Betaine-homocysteine methyl transferase (BHMT) catalyzes the synthesis of methionine from betaine and homocysteine (Hcy), utilizing a zinc ion to activate Hcy. BHMT is a key liver enzyme that is important for homocysteine homeostasis. X-ray structures of human BHMT in its oxidized (Zn-free) and reduced (Zn-replete) forms, the latter in complex with the bisubstrate analog, S(delta-carboxybutyl)-L-homocysteine, were determined at resolutions of 2.15 A and 2.05 A. BHMT is a (beta/alpha)(8) barrel that is distorted to construct the substrate and metal binding sites. The zinc binding sequences G-V/L-N-C and G-G-C-C are at the C termini of strands beta6 and beta8. Oxidation to the Cys217-Cys299 disulfide and expulsion of Zn are accompanied by local rearrangements. The structures identify Hcy binding fingerprints and provide a prototype for the homocysteine S-methyltransferase family.
- Biophysics Research Division and Department of Biological Chemistry, University of Michigan, Ann Arbor, MI 48109, USA.
Organizational Affiliation: