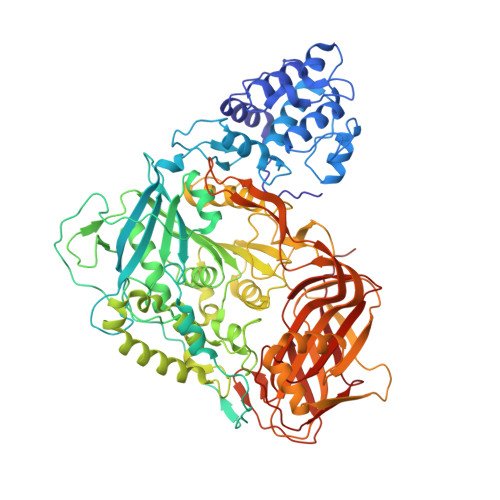
The Structural Basis for Catalysis and Specificity of the X-prolyl dipepdidyl aminopeptidase from Lactococcus lactis
Rigolet, P., Mechin, I., Delage, M.M., Chich, J.F.(2002) Structure 10: 1383-1394
- PubMed: 12377124 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00851-1
- Primary Citation Related Structures:
1LNS - PubMed Abstract:
The X-prolyl dipeptidyl aminopeptidase (X-PDAP) from Lactococcus lactis is a dimeric enzyme catalyzing the removal of Xaa-Pro dipeptides from the N terminus of peptides. The structure of the enzyme was solved at 2.2 A resolution and provides a model for the peptidase family S15. Each monomer is composed of four domains. The larger one presents an alpha/beta hydrolase fold and comprises the active site serine. The specificity pocket is mainly built by residues from a small helical domain which is, together with the N-terminal domain, essential for dimerization. A C-terminal moiety probably plays a role in the tropism of X-PDAP toward the cellular membrane. These results give new insights for further exploration of the role of the enzymes of the SC clan.
- L.U.R.E. Centre Universitaire Paris-Sud, Bât. 209D-BP 34, F-91898 Orsay Cedex, France. rigolet@lure.u-psud.fr
Organizational Affiliation: