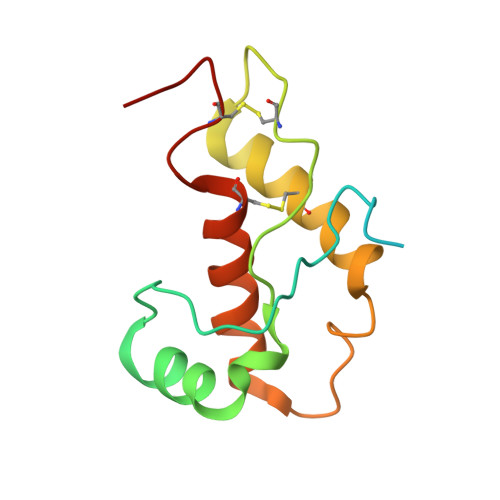
NMR Structure of the Human Doppel Protein
Luhrs, T., Riek, R., Guntert, P., Wuthrich, K.(2003) J Mol Biology 326: 1549-1557
- PubMed: 12595265 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)01471-7
- Primary Citation Related Structures:
1LG4 - PubMed Abstract:
The NMR structure of the recombinant human doppel protein, hDpl(24-152), contains a flexibly disordered "tail" comprising residues 24-51, and a globular domain extending from residues 52 to 149 for which a detailed structure was obtained. The globular domain contains four alpha-helices comprising residues 72-80 (alpha1), 101-115 (alpha2(a)), 117-121 (alpha2(b)), and 127-141 (alpha3), and a short two-stranded anti-parallel beta-sheet comprising residues 58-60 (beta1) and 88-90 (beta2). The fold of the hDpl globular domain thus coincides nearly identically with the structure of the murine Dpl protein. There are close similarities with the human prion protein (hPrP) but, similar to the situation with the corresponding murine proteins, hDpl shows marked local differences when compared to hPrP: the beta-sheet is flipped by 180 degrees with respect to the molecular scaffold formed by the four helices, and the beta1-strand is shifted by two residues toward the C terminus. A large solvent-accessible hydrophobic cleft is formed on the protein surface between beta2 and alpha3, which has no counterpart in hPrP. The helix alpha2 of hPrP is replaced by two shorter helices, alpha2(a) and alpha2(b). The helix alpha3 is shortened by more than two turns when compared with alpha3 of hPrP, which is enforced by the positioning of the second disulfide bond in hDpl. The C-terminal peptide segment 144-149 folds back onto the loop connecting beta2 and alpha2. All but four of the 20 conserved residues in the globular domains of hPrP and hDpl appear to have a structural role in maintaining a PrP-type fold. The conservation of R76, E96, N110 and R134 in hDpl, corresponding to R148, E168, N183 and R208 in hPrP suggests that these amino acid residues might have essential roles in the so far unknown functions of PrP and Dpl in healthy organisms.
- Institut für Molekularbiologie und Biophysik, Eidgenössische Technische Hochschule Zürich, CH-8093, Zürich, Switzerland. luehrs@salk.edu
Organizational Affiliation: