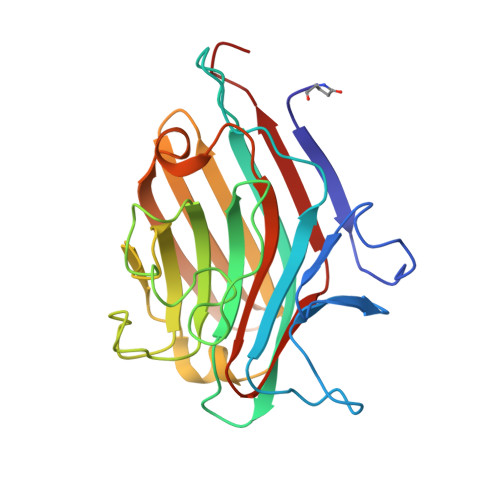
Structures of the lectin IV of Griffonia simplicifolia and its complex with the Lewis b human blood group determinant at 2.0 A resolution.
Delbaere, L.T., Vandonselaar, M., Prasad, L., Quail, J.W., Wilson, K.S., Dauter, Z.(1993) J Mol Biology 230: 950-965
- PubMed: 8478943 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1212
- Primary Citation Related Structures:
1GSL, 1LEC, 1LED - PubMed Abstract:
The structures of the fourth lectin isolated from Griffonia simplicifolia (GS4) and its complex with the methyl-glycoside of the Lewis b human blood group determinant (Le(b)-OMe) are reported at high resolution. The native GS4 crystal is isomorphous with the complexed GS4 crystal. The space group is P4(2)2(1)2 with unit cell dimensions a = 78.9 A, c = 89.1 A with one subunit of the lectin (bound to 1 Le(b)-OMe in the complex) in the crystallographic asymmetric unit. The native GS4 structure was solved by the molecular replacement technique and least-squares refined (PROLSQ and X-PLOR). The orientation of the Le(b)-OMe tetrasaccharide in the complex was established from a 2.8 A difference map with coefficients (Fcomplex--Fnative) and calculated phase angles from the native model. Both the final native and complex GS4 models consist of 1904 protein non-hydrogen atoms, one sulfate ion, one Ca ion, one Mn ion and three covalently-bound sugar residues N-linked to Asn18. In addition, the complex model has 47 Le(b)-OMe non-hydrogen atoms. The two structures have 135 water molecules in common in addition to eight and nine unique water molecules in the native and complex structures, respectively. The root-mean-square deviations from ideal bond distances and angles are 0.016 A, 3.2 degrees and 0.016 A, 3.0 degrees, for the native and complexed GS4, respectively. The R index for all unique data from 8 to 2.0 A is 0.187 for the native (19,204 reflections) and 0.181 for the complex (19,212 reflections). The tertiary structure of each subunit is similar to that of other leguminous lectins but the quaternary structure of the molecular dimer is different from that of any other lectin reported to date. The co-ordination about the Ca ion is pentagonal bipyramidal (with 1 long Ca(2+)-oxygen bond) and the co-ordination about the Mn ion is octahedral. Two conserved residues (Asp149 and Ser155) appear to be important because they are hydrogen-bonded to each other and to groups that co-ordinate the Mn ion. There are three cis-peptides in the polypeptide chain; two involve non-proline residues, one of which is homologous with other leguminous lectins and the other is unique to GS4. The two non-proline cis-peptides are located in the carbohydrate-binding site and are important for the specificity of the lectin. The molecular recognition of Le(b)-OMe by GS4 involves both polar and extensive non-polar interactions.(ABSTRACT TRUNCATED AT 400 WORDS)
- Department of Biochemistry, University of Saskatchewan, Saskatoon, Canada.
Organizational Affiliation: