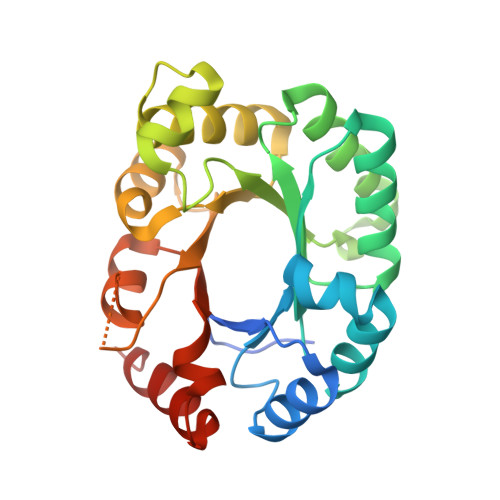
Substrate binding induces domain movements in orotidine 5'-monophosphate decarboxylase
Harris, P., Poulsen, J.C., Jensen, K.F., Larsen, S.(2002) J Mol Biology 18: 1019-1029
- PubMed: 12054799 Search on PubMed
- DOI: https://doi.org/10.1016/S0022-2836(02)00200-0
- Primary Citation Related Structures:
1L2U - PubMed Abstract:
Orotidine 5'-monophosphate decarboxylase (ODCase) catalyses the decarboxylation of orotidine 5'-monophosphate to uridine 5'-monophosphate (UMP). We have earlier determined the structure of ODCase from Escherichia coli complexed with the inhibitor 1-(5'-phospho-beta-d-ribofuranosyl)barbituric acid (BMP); here we present the 2.5 A structure of the uncomplexed apo enzyme, determined from twinned crystals. A structural analysis and comparison of the two structures of the E. coli enzyme show that binding of the inhibitor is accompanied by significant domain movements of approximately 12 degrees around a hinge that crosses the active site. Hence, the ODCase dimer, which contains two active sites, may be divided in three domains: a central domain that is fixed, and two lids which independently move 12 degrees upon binding. Corresponding analyses, presented herein, of the two Saccharomyces cerevisiae ODCase structures (with and without BMP) and the Methanobacterium thermoautotrophicum ODCase structures (with and without 6-aza UMP) show very similar, but somewhat smaller domain movements. The domain movements seem to be initiated by the phosphoryl binding to the enzyme and can explain why the binding of the phosphoryl group is essential for the catalytic function.
- Centre for Crystallographic Studies, Department of Chemistry, University of Copenhagen, Universitetsparken 5, Denmark.
Organizational Affiliation: