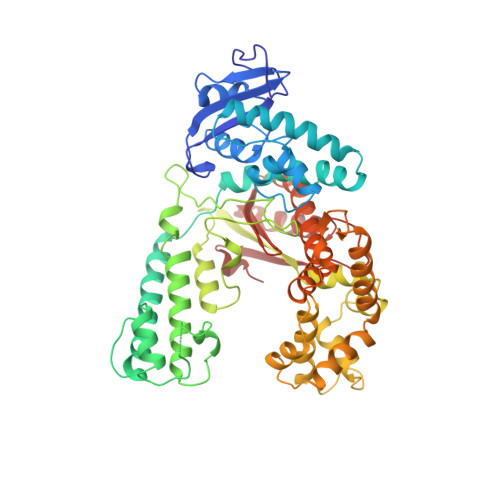
Crystal structure of the large fragment of Thermus aquaticus DNA polymerase I at 2.5-A resolution: structural basis for thermostability.
Korolev, S., Nayal, M., Barnes, W.M., Di Cera, E., Waksman, G.(1995) Proc Natl Acad Sci U S A 92: 9264-9268
- PubMed: 7568114 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.92.20.9264
- Primary Citation Related Structures:
1KTQ - PubMed Abstract:
The crystal structure of the large fragment of the Thermus aquaticus DNA polymerase (Klentaq1), determined at 2.5-A resolution, demonstrates a compact two-domain architecture. The C-terminal domain is identical in fold to the equivalent region of the Klenow fragment of Escherichia coli DNA polymerase I (Klenow pol I). Although the N-terminal domain of Klentaq1 differs greatly in sequence from its counterpart in Klenow pol I, it has clearly evolved from a common ancestor. The structure of Klentaq1 reveals the strategy utilized by this protein to maintain activity at high temperatures and provides the structural basis for future improvements of the enzyme.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, MO 63110, USA.
Organizational Affiliation: