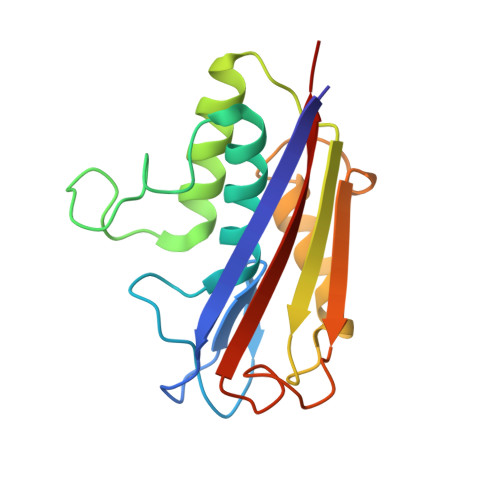
Structure and mechanism of 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase. An enzyme in the mevalonate-independent isoprenoid biosynthetic pathway.
Richard, S.B., Ferrer, J.L., Bowman, M.E., Lillo, A.M., Tetzlaff, C.N., Cane, D.E., Noel, J.P.(2002) J Biological Chem 277: 8667-8672
- PubMed: 11786530 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.C100739200
- Primary Citation Related Structures:
1KNJ, 1KNK - PubMed Abstract:
The enzyme 2-C-methyl-D-erythritol 2,4-cyclodiphosphate (MECDP) synthase catalyzes the conversion of 4-diphosphocytidyl-2-C-methyl-D-erythritol 2-phosphate (CDP-ME2P) to MECDP, a highly unusual cyclodiphosphate-containing intermediate on the mevalonate-independent pathway to isopentenyl diphosphate and dimethylallyl diphosphate. We now report two x-ray crystal structures of MECDP synthase refined to 2.8-A resolution. The first structure contains a bound Mn(2+) cation, and the second structure contains CMP, MECDP, and Mn(2+). The protein adopts a homotrimeric quaternary structure built around a central hydrophobic cavity and three externally facing active sites. Each of these active sites is located between two adjacent monomers. A tetrahedrally arranged transition metal binding site, potentially occupied by Mn(2+), sits at the base of the active site cleft. A phosphate oxygen of MECDP and the side chains of Asp(8), His(10), and His(42) occupy the metal ion coordination sphere. These structures reveal for the first time the structural determinants underlying substrate, product, and Mn(2+) recognition and the likely catalytic mechanism accompanying the biosynthesis of the cyclodiphosphate-containing isoprenoid precursor, MECDP.
- Structural Biology Laboratory, The Salk Institute for Biological Studies, La Jolla, California 92037, USA.
Organizational Affiliation: