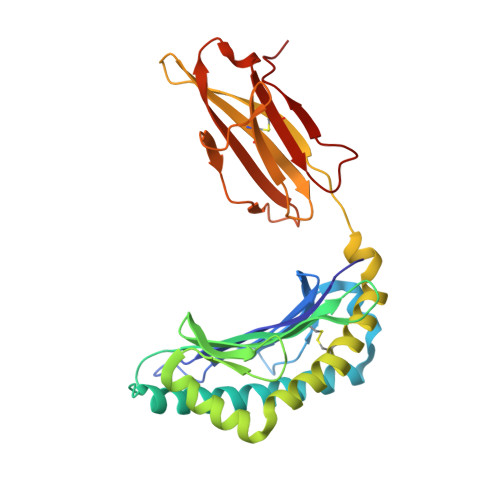
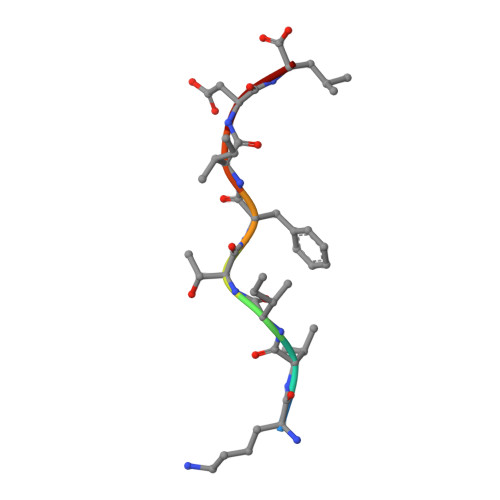
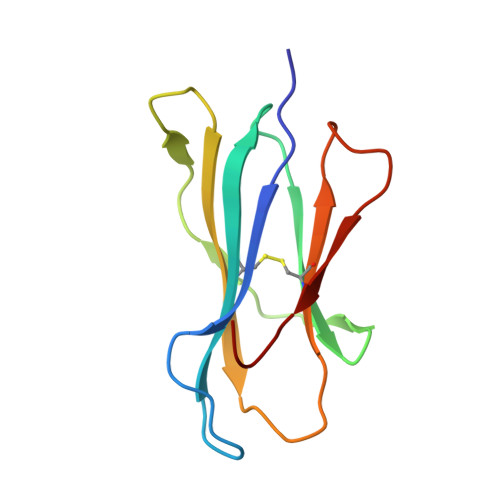
A T cell receptor CDR3beta loop undergoes conformational changes of unprecedented magnitude upon binding to a peptide/MHC class I complex.
Reiser, J.B., Gregoire, C., Darnault, C., Mosser, T., Guimezanes, A., Schmitt-Verhulst, A.M., Fontecilla-Camps, J.C., Mazza, G., Malissen, B., Housset, D.(2002) Immunity 16: 345-354
- PubMed: 11911820 Search on PubMed
- DOI: https://doi.org/10.1016/s1074-7613(02)00288-1
- Primary Citation Related Structures:
1KJ2, 1KJ3 - PubMed Abstract:
The elongated complementary-determining region (CDR) 3beta found in the unliganded KB5-C20 TCR protrudes from the antigen binding site and prevents its docking onto the peptide/MHC (pMHC) surface according to a canonical diagonal orientation. We now present the crystal structure of a complex involving the KB5-C20 TCR and an octapeptide bound to the allogeneic H-2K(b) MHC class I molecule. This structure reveals how a tremendously large CDR3beta conformational change allows the KB5-C20 TCR to adapt to the rather constrained pMHC surface and achieve a diagonal docking mode. This extreme case of induced fit also shows that TCR plasticity is primarily restricted to CDR3 loops and does not propagate away from the antigen binding site.
- Laboratoire de Cristallographie et Cristallogénèse des Protéines, Institut de Biologie Structurale J.-P. Ebel, CEA-CNRS-UJF, 41 rue Jules Horowitz, F-38027 Grenoble Cedex 1, France.
Organizational Affiliation: