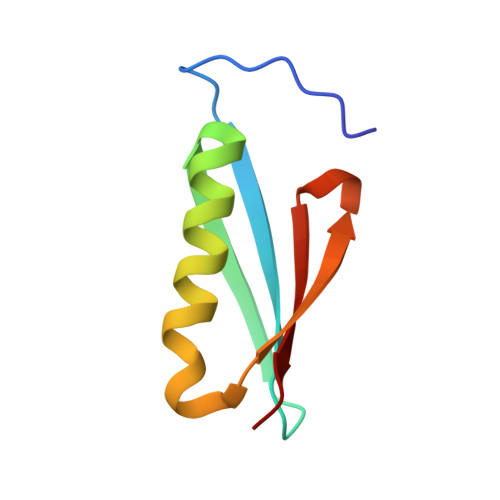
Single-site mutations induce 3D domain swapping in the B1 domain of protein L from Peptostreptococcus magnus.
O'Neill, J.W., Kim, D.E., Johnsen, K., Baker, D., Zhang, K.Y.(2001) Structure 9: 1017-1027
- PubMed: 11709166 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00667-0
- Primary Citation Related Structures:
1K50, 1K51, 1K52, 1K53 - PubMed Abstract:
Thermodynamic and kinetic studies of the Protein L B1 domain (Ppl) suggest a folding pathway in which, during the folding transition, the first beta hairpin is formed while the second beta hairpin and the alpha helix are largely unstructured. The same mutations in the two beta turns have opposite effects on the folding and unfolding rates. Three of the four residues composing the second beta turn in Ppl have consecutive positive phi angles, indicating strain in the second beta turn. We have determined the crystal structures of the beta turn mutants G55A, K54G, and G15A, as well as a core mutant, V49A, in order to investigate how backbone strain affects the overall structure of Ppl. Perturbation of the hydrophobic interactions at the closed interface by the V49A mutation triggered the domain swapping of the C-terminal beta strand that relieved the strain in the second beta turn. Interestingly, the asymmetric unit of V49A contains two monomers and one domain-swapped dimer. The G55A mutation escalated the strain in the second beta turn, and this increased strain shifted the equilibrium toward the domain-swapped dimer. The K54G structure revealed that the increased stability is due to the reduction of strain in the second beta turn, while the G15A structure showed that increased strain alone is insufficient to trigger domain swapping. Domain swapping in Ppl is determined by the balance of two opposing components of the free energy. One is the strain in the second beta turn that favors the dimer, and the other is the entropic cost of dimer formation that favors the monomer. A single-site mutation can disrupt this balance and trigger domain swapping.
- Division of Basic Sciences, Fred Hutchinson Cancer Research Center, 1100 Fairview Avenue North, Seattle, WA 98109, USA.
Organizational Affiliation: