Structural identification of a bacterial quorum-sensing signal containing boron.
Chen, X., Schauder, S., Potier, N., Van Dorsselaer, A., Pelczer, I., Bassler, B.L., Hughson, F.M.(2002) Nature 415: 545-549
- PubMed: 11823863 Search on PubMed
- DOI: https://doi.org/10.1038/415545a
- Primary Citation Related Structures:
1JX6 - PubMed Abstract:
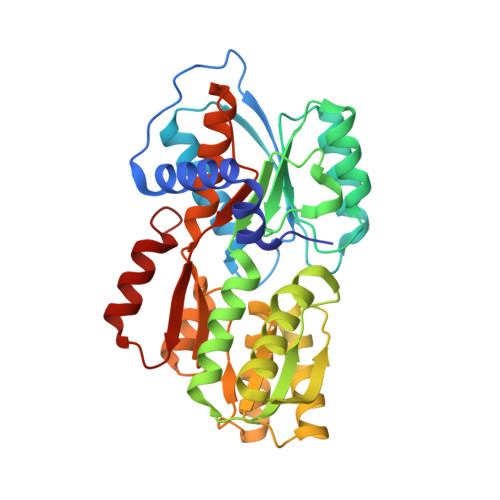
Cell-cell communication in bacteria is accomplished through the exchange of extracellular signalling molecules called autoinducers. This process, termed quorum sensing, allows bacterial populations to coordinate gene expression. Community cooperation probably enhances the effectiveness of processes such as bioluminescence, virulence factor expression, antibiotic production and biofilm development. Unlike other autoinducers, which are specific to a particular species of bacteria, a recently discovered autoinducer (AI-2) is produced by a large number of bacterial species. AI-2 has been proposed to serve as a 'universal' signal for inter-species communication. The chemical identity of AI-2 has, however, proved elusive. Here we present the crystal structure of an AI-2 sensor protein, LuxP, in a complex with autoinducer. The bound ligand is a furanosyl borate diester that bears no resemblance to previously characterized autoinducers. Our findings suggest that addition of naturally occurring borate to an AI-2 precursor generates active AI-2. Furthermore, they indicate a potential biological role for boron, an element required by a number of organisms but for unknown reasons.
- Department of Molecular Biology, Princeton University, Princeton, New Jersey 08544-1014, USA.
Organizational Affiliation: