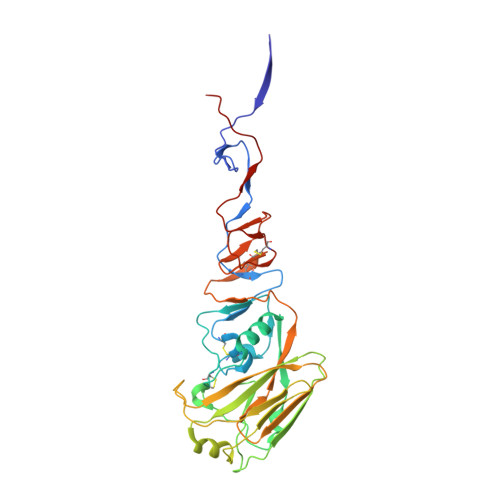
H5 avian and H9 swine influenza virus haemagglutinin structures: possible origin of influenza subtypes.
Ha, Y., Stevens, D.J., Skehel, J.J., Wiley, D.C.(2002) EMBO J 21: 865-875
- PubMed: 11867515 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/21.5.865
- Primary Citation Related Structures:
1JSD, 1JSM - PubMed Abstract:
There are 15 subtypes of influenza A virus (H1-H15), all of which are found in avian species. Three caused pandemics in the last century: H1 in 1918 (and 1977), H2 in 1957 and H3 in 1968. In 1997, an H5 avian virus and in 1999 an H9 virus caused outbreaks of respiratory disease in Hong Kong. We have determined the three-dimensional structures of the haemagglutinins (HAs) from H5 avian and H9 swine viruses closely related to the viruses isolated from humans in Hong Kong. We have compared them with known structures of the H3 HA from the virus that caused the 1968 H3 pandemic and of the HA--esterase--fusion (HEF) glycoprotein from an influenza C virus. Structure and sequence comparisons suggest that HA subtypes may have originated by diversification of properties that affected the metastability of HAs required for their membrane fusion activities in viral infection.
- Department of Molecular and Cellular Biology, Harvard University, 7 Divinity Avenue, Cambridge, MA 02138, USA.
Organizational Affiliation: